Key Points
-
Staphylococcus aureus evades innate and adaptive immune responses to cause localized or systemic infections in humans. Because the development of protective immunity is prevented, S. aureus infections reoccur even with antibiotic or surgical therapy.
-
Mechanisms have been revealed whereby products secreted by S. aureus interfere with neutrophil chemotaxis, complement activation, opsonization and phagocytic killing of bacteria. Immune evasion determinants can interact with host factors from humans, but not with counterparts from other vertebrates, which presents a challenge for the development of animal models.
-
The hallmark of S. aureus is the secretion of coagulases that associate with prothrombin to generate fibrin clots. Through the fibrinogen and fibrin binding attributes of staphylococcal surface proteins, the pathogen shields itself from host phagocytes, which is a prerequisite for abscess lesion formation in infected tissues. Macrophage access to these lesions is restricted by staphylococcal-induced degradation of neutrophil extracellular traps (NETs).
-
Staphylococcal protein A (SpA) crosslinks B cell receptors and triggers proliferative expansion of VH3+ B cells and the secretion of antibodies that fail to recognize S. aureus antigens. SpA blocks host antibody responses that are required for the establishment of protective immunity.
-
T cell superantigens crosslink major histocompatibility class II molecules of antigen-presenting cells with the T cell receptor, promoting lymphocyte proliferation, anergy and the release of cytokines (a cytokine storm). Superantigens vary between S. aureus strains and activate distinct subsets of Vβ chain T cell receptors, endowing staphylococcal isolates with unique T cell-avoidance attributes.
-
Genome sequencing of S. aureus isolates from humans or domesticated animals revealed that only some immune evasion genes are conserved among all strains. Even conserved genes display sequence polymorphisms, which presents a formidable challenge for the design of S. aureus vaccines. Panoplies of immune evasion factors endow staphylococcal strains with unique virulence attributes and with the ability for epidemic spread.
Abstract
Staphylococcus aureus, a bacterial commensal of the human nares and skin, is a frequent cause of soft tissue and bloodstream infections. A hallmark of staphylococcal infections is their frequent recurrence, even when treated with antibiotics and surgical intervention, which demonstrates the bacterium's ability to manipulate innate and adaptive immune responses. In this Review, we highlight how S. aureus virulence factors inhibit complement activation, block and destroy phagocytic cells and modify host B cell and T cell responses, and we discuss how these insights might be useful for the development of novel therapies against infections with antibiotic resistant strains such as methicillin-resistant S. aureus.
Similar content being viewed by others
Main
Approximately 30% of the human population is continuously colonized with Staphylococcus aureus, whereas some individuals are hosts for intermittent colonization1. S. aureus typically resides in the nares but is also found on the skin and in the gastrointestinal tract. Although colonization is not a prerequisite for staphylococcal disease, colonized individuals more frequently acquire infections1. Skin and soft-tissue infections (SSTIs) are the most frequent disease form of S. aureus, and these infections can progress to bacteraemia and invasive disease (that is, bloodstream infection, endocarditis or sepsis2). In addition, S. aureus can cause pneumonia, osteomyelitis, infectious arthritis, abscesses in many organ tissues and infections of surgical wounds or prosthetic materials2. Annual attack rates for S. aureus disease range between 1–3% and vary with age, ethnicity and geographical location of human populations2. At elevated risk for staphylococcal infection are low-birth-weight infants, children, the elderly and patients with indwelling catheters, endotracheal intubation, medical implantation of foreign bodies, trauma, surgical procedures, haemodialysis, diabetes or immunosuppressive or cancer therapy2. A key feature of S. aureus disease is its recurrence, which occurs for 8–33% of cases of SSTI and bloodstream infections3. Prior disease does not elicit protection against subsequent S. aureus infection2.
Neutrophils play a central part in protecting humans against S. aureus infection. Staphylococcal entry and replication in host tissues leads to the release of bacterial products (formyl-peptides, lipoproteins or peptidoglycan) and to damaged tissues that produce inflammatory signals (that is, chemoattractants and cytokines4). Staphylococcal products are detected by immune cells via Toll-like receptors (TLRs) and G protein-coupled receptors, whereas cytokines activate cognate immune receptors. Neutrophils answer this call, extravasate from blood vessels, and migrate towards the site of infection to phagocytose and kill bacteria or to immobilize and damage the pathogen through NETosis — the release of neutrophil extracellular traps (NETs) comprising DNA and antimicrobial peptides4. The importance of neutrophils in controlling S. aureus infection has been documented through the study of immune defects. Mutations in genes encoding NADPH oxidase, the enzyme generating bactericidal superoxide in phagocytes, cause chronic granulomatous disease (CGD), which is associated with defects in phagocytic killing of S. aureus and frequent infection5. Individuals with inborn errors of signal transducer and activator of transcription 1 (STAT1) or STAT3 signalling of immune cells have perturbed interleukin-17 (IL-17) cytokine pathways, which diminishes mucocutaneous immunity and promotes S. aureus infection6. IL-17-dependent T cell signalling is a key activator of neutrophils and of anti-staphylococcal defences7. Finally, cancer patients with diminished blood neutrophil counts are highly susceptible to S. aureus infection8.
Nevertheless, the vast majority of S. aureus disease occurs in immune-competent individuals without defects in phagocyte function. To achieve this, S. aureus deploys an arsenal of immune-evasive strategies that together prevent phagocytosis and killing by neutrophils. Furthermore, the ability of the pathogen to cause recurrent disease implies the presence of mechanisms that effectively block the development of adaptive immune responses. Here, we review recent work on the immune evasive attributes of S. aureus, including the subversion of the innate and adaptive immune systems and the killing of immune cells, along with epidemiological features of the corresponding genes. We also discuss how the characterization of bacterial immune evasive factors can have translational effects in the therapy of autoimmune diseases or the development of vaccines and immunotherapeutics against S. aureus infection.
Subversion of innate immune responses
Neutrophil extravasation and chemotaxis. Pro-inflammatory signals promote neutrophil adhesion and extravasation across capillary endothelia, relying on reciprocal interactions between endothelial receptors (such as P-selectin, E selectin, intercellular adhesion molecule 1 (ICAM1) and hyaluronan) and ligands on neutrophil surfaces (such as P-selectin glycoprotein ligand 1 (PSGL1), lymphocyte function-associated antigen 1 (LFA1; also known as αLβ2 integrin), αMβ2 integrin and CD44)9. Although neutrophils function to migrate towards bacterial invaders, S. aureus can interfere with neutrophil extravasation and chemotaxis through the secretion of staphylococcal superantigen-like proteins (SSLs), phenol-soluble modulins (PSMs), chemotaxis inhibitory protein of S. aureus (CHIPS), formyl peptide receptor-like 1 inhibitor (FLIPr) and its homologue FLIPr-like (FLIPrL).
SSLs are a family of secreted proteins with structural homology to staphylococcal superantigens10,11,12. The ssl genes are arranged as tandem repeats in genomic island-α (GIα; for ssl1–ssl11) and in the immune evasion cluster 2 (IEC2; for ssl12–ssl14) on the bacterial chromosome13. GIα-encoded ssl genes vary between lineages as does the coding sequence of individual ssl genes; the number of different alleles ranges from 1 to 13 and most alleles are uniquely associated with specific S. aureus lineages13. ssl1, ssl2, ssl3, ssl11, ssl12, ssl13 and ssl14 are found in all S. aureus isolates13 (Box 1). Purified, recombinant SSL5 and SSL11 bind PSGL1 on leukocytes and, when assayed in vitro, interfere with the binding of neutrophils to P-selectin and neutrophil adhesion and rolling14,15 (Fig. 1a). SSL5 also interferes with chemokine- and anaphylatoxin-mediated activation of neutrophils by binding to the glycosylated amino termini of G protein-coupled receptors14,16. Moreover, SSL5 has been shown to activate platelets and support their adhesion involving platelet surface receptors GPIbα and GPVI17,18. Intravenous administration of SSL5 caused intravascular platelet-rich thrombi and increased bleeding of C57Bl/6 mice19. Other work demonstrated SSL5-mediated inactivation of leukocyte matrix metalloproteinase 9 (Ref. 20). The affinity of SSL5 for different host factors is mediated via its glycan binding pocket, an attribute that is shared by other members of the SSL family21. SSL3 binds to TLR2 and blocks immune cell recognition of staphylococcal lipoproteins and peptidoglycan via TLR1–TLR2 and TLR2–TLR6 heterodimers22, and SSL10 blocks C-X-C chemokine receptor 4 (CXCR4)-mediated responses on lymphocytes, interfering with the chemoattraction of neutrophils4 (Fig. 1a). Recent work suggests that SelX, a staphylococcal enterotoxin (superantigen)-like protein, also binds glycosylated PSGL1, and that SSL6 binds to CD47 (also known as integrin-associated protein), a common receptor on most host tissues that promotes migration, anti-phagocytosis and proliferation23. Thus, SSLs presumably associate with a wide range of glycoproteins on leukocytes and/or platelets to implement immune-evasive attributes. Most SSLs display species specificity for human but not animal host factors, and SSL-mediated contributions towards S. aureus pathogenesis cannot be measured in animal experiments. S. aureus also inhibits leukocyte migration via the extracellular adherence protein (Eap). Eap is composed of four β-grasp-like domains and associates with ICAM1 to inhibit leukocyte migration24. The eap gene is located in the eap–hlb locus, the attachment site for hlb-converting phages carrying the IEC1 gene cluster13 (Box 1).
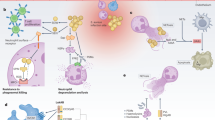
a | Neutrophil extravasation and chemotaxis is inhibited by Staphylococcus aureus through the secretion of staphylococcal superantigen-like (SSL) molecules. SSL3 inhibits Toll-like receptor (TLR) heterodimers, SSL5, SSL11 and SelX inhibit PSGL1 signalling (SSL11 is not shown), and SSL6 inhibits the G protein-coupled receptor CD47. Other secreted proteins include chemotaxis inhibitory protein of S. aureus (CHIPS), which inhibits the complement receptor C5aR and formyl-peptide receptor 1 (FPR1) and FPR2, formyl peptide receptor-like 1 inhibitor (FLIPr) and FLIPr-like (FLIPrL), which inhibit FPR1 and FPR2, and staphopain, which inhibits signalling from the chemokine receptor C-X-C chemokine receptor (CXCR2). b | Complement activation and phagocytosis of staphylococci are blocked through the secretion of inhibitory factors that interfere with opsonization. Collagen adhesin (Cna) blocks the association of complement factor C1q bound to immunoglobulin with complement receptor C1r. Staphylococcal protein A (SpA) and staphylococcal binder of immunoglobulin (Sbi) binding to immunoglobulin blocks its association with C1q. Sbi, SpA, SSL7 and SSL10 sequester immunoglobulins to block their ability to promote complement activation. Sbi (when associated with the host factors C3d and factor H (fH)) and SSL7 also inactivate the complement factors C3 and C5, respectively. Sak associates with plasminogen (PLG) and activates the zymogen to cleave complement factor C3b and immunoglobulin. Extracellular complement-binding protein (Ecb), extracellular fibrinogen-binding protein (Efb), staphylococcal complement inhibitor (SCIN) and extracellular adherence protein (Eap) inhibit C3 convertases, and aureolysin (Aur) cleaves the complement factor C3, which compromises opsonization because the cleavage product C3b is degraded by a complex of the host proteins fI and fH. c | S. aureus inhibits neutrophil-mediated killing of phagocytosed bacteria by expressing several enzymes and inhibitors. The adenosine-synthesizing enzyme AdsA enables the inhibition of granulation via adenosine receptor (AdoR) signalling. Staphyloxanthin, superoxide dismutase A (SodA) and SodM, the catalase KatG and alkylhydroperoxide reductase (AhpC) are antioxidants that reduce oxidative stress caused by phagosomal reactive oxygen species (ROS) and H2O2 generation. Aureolysin (Aur) cleaves antimicrobial peptides and DltA–DltD promote d-alanyl esterification of teichoic acids to protect staphylococci from antimicrobial peptides. MprF modifies phosphatidylglycerol with alanine or lysine, another mechanism to protect staphylococci against antimicrobial peptides. l-lactate dehydrogenase (Ldh) and flavohaemoglobin (Hmp) inhibit nitrosative stress, Eap and its homologues EapH1 and EapH2 inhibit neutrophil serine proteases, and OatA O-acetylates peptidoglycan, which prevents its lysozymal degradation. d | Secreted β-barrel pore forming toxins (β-PFTs) bind specific receptors on immune cells to impair immune cell functions or promote cell lysis. These β-PFTs include leukocidin ED (LukED) (which binds to neutrophils, T cells and macrophages), γ-haemolysin AB (HlgAB) (which binds to neutrophils, macrophages and red blood cells), HlgCB and Panton–Valentine leukocidin (PVL) (which bind to neutrophils and macrophages), and LukAB and α-haemolysin (Hla) (which bind to neutrophils). Phenol-soluble modulin-α (PSMα), which is another factor secreted by S. aureus (but not a β-PFT), can also lyse leukocytes.
PSMs are a family of short formyl-peptides that are secreted via an ABC transporter and interfere with the physiological functions of immune cells, specifically neutrophils25. PSMα1–PSMα4 and PSMβ1, and PSMβ2, whether formylated or not, activate formyl-peptide receptor 2 (FPR2) on human and mouse neutrophils and stimulate cytokine release26. Of note, PSMα1–PSMα4 and PSMβ1 and PSMβ2 are neutralized by binding to serum lipoproteins, and reactive oxygen species (ROS) of activated neutrophils are reported to neutralize PSM signalling27. Recent work demonstrated that PSMα3N22Y, a variant with diminished FPR2-binding activity that is secreted by the clonal complex 30 (CC30) methicillin-resistant S. aureus (MRSA) lineage, is associated with diminished FPR2 signalling and diminished cytotoxicity, while enhancing bacterial replication and the establishment of abscess lesions in renal tissues28. The activation of FPRs via the formyl moiety of PSMs and via their direct binding to FPR2 stimulates chemotaxis. Thus, a key attribute of formylated PSMs seems to be the stimulation of neutrophil chemotaxis via FPRs.
S. aureus also counters neutrophil chemotaxis, which occurs via the secretion of CHIPS, FLIPr and FLIPrL4 (Fig. 1a). CHIPS is encoded by the chp gene in IEC1, which is carried on hlb-converting phages29. chp is found in most human S. aureus isolates but not in livestock-associated strains13. Secreted CHIPS binds to human formyl-peptide receptor 1 (FPR1) and human C5aR, the complement receptor of neutrophils, but not to mouse FPR1 or C5aR30,31. FLIPr and FLIPrL are encoded by genes in the IEC2 locus; at least 9 different alleles are known for these genes, which are found in many, but not all, human S. aureus isolates13 (Box 1). FLIPrL inhibits FPR1 signalling, whereas FLIPr and FLIPrL bind to FPR2 and inhibit its receptor signalling function32,33. Finally, staphopain (ScpA), a secreted cysteine protease, cleaves CXCR2-binding chemokines to block neutrophil migration towards staphylococci34 (Fig. 1a). The lifestyle of S. aureus — invasion of host tissues, replication in abscess lesions and dissemination upon purulent drainage of lesions — requires recruitment of immune cells to the site of infection35,36. However, staphylococcal products manipulate infiltrating immune cells to limit their capacity for chemotaxis, phagocytosis and bacterial killing, thereby ensuring the successful outcome of infection.
Complement activation and phagocytosis. Complement, a key component of innate host defences, is composed of >30 proteins that have broad functions in host defence against microorganisms, inflammation, haemostasis and wound repair. Complement is activated by any one of three routes: the classical pathway (which entails antibody and C1q deposition on the staphylococcal surface), the lectin pathway (which involves the association of mannose-binding lectin (MBL)–MBL-associated serine protease (MASP) complexes with staphylococcal carbohydrates) and the alternative pathway (which involves spontaneous breakdown of the complement protein C3 in the serum). The three pathways converge in the assembly of a C3 convertase (C4b2a for the classical and lectin pathways, and C3bBb for the alternative pathway). C3 convertase cleaves C3 into C3a and C3b; C3b becomes covalently linked to the staphylococcal surface (known as opsonization), and C3a is released as a chemoattractant for phagocytes37. At high local concentrations of C3b, the C5 convertase cleaves C5 into C5a, another chemoattractant, and C5b, the surface deposition of which promotes membrane attack complex (MAC) formation37. The MAC is effective at killing Gram-negative bacteria; however, it is not effective against S. aureus, which has a thick peptidoglycan layer that prevents access to the bacterial membrane38.
In addition to the thick peptidoglycan layer, many clinical S. aureus strains express genes for the production of one of two types of capsular polysaccharide, type 5 or type 8 (Ref. 39). In vitro phagocytosis assays suggested that capsule expression protects staphylococci from neutrophil phagocytosis in the presence of opsonins, and that capsule expression contributes to the pathogenesis of S. aureus infection in mice39. Capsule-induced protection from phagocytosis may be strain specific, as capsule mutations in other S. aureus isolates do not affect the pathogenesis of bloodstream infections in mice36. USA300, the current pandemic clone of community-acquired MRSA infections, carries a mutation that abrogates capsule expression40 (Table 1).
S. aureus secretes several proteins that interfere with the deposition of complement on the bacterial surface (Fig. 1b). Aureolysin, a secreted zinc-dependent metalloproteinase, cleaves C3 to generate functionally active C3a and C3b. Complement factor I (fI) and factor H (fH) degrade or bind C3b, which prevents its accumulation on the staphylococcal surface41. The aureolysin gene, aur, is polymorphic and specific alleles are associated with different S. aureus lineages. An in vivo phenotype for S. aureus aur mutants has not yet been described.
Staphylococcal complement inhibitor (SCIN) associates with and inhibits C3 convertase (C3bBb), thereby preventing the production of C3a, C3b and C5a and interfering with complement activation42 (Fig. 1b). The structural gene for SCIN, scn, is also located on hlb-converting phages together with chp and sak (which encodes staphylokinase, see below). Two polymorphic homologues of SCIN, designated SCIN-B and SCIN-C, are encoded by genes in the IEC2 locus. scn, scnB and scnC are found in many, but not all, human clinical isolates (Box 1); SCIN, SCIN-B and SCIN-C associate with C3 convertase from humans but not with that of other vertebrates43.
The genes encoding extracellular fibrinogen-binding protein (Efb) and its homologue, extracellular complement-binding protein (Ecb), are also located on IEC2. Both Efb and Ecb bind to C3d (a cleavage product of C3b that activates innate and adaptive responses by binding to complement receptor 2 (CR2)) and inhibit C3bBb and the C5 convertases43,44 (Fig. 1b). Ecb associates with both fH and C3b to facilitate the complement inhibitory attributes of fH45. Efb also binds fibrinogen and prevents fibrinogen interaction with αMβ2, an integrin on neutrophils that activates pro-inflammatory responses, as well as fibrinogen-mediated platelet activation46,47. Efb and Ecb inhibitory activities have been observed for human and mouse convertases and fibrinogen. In the mouse intravenous challenge model, the S. aureus ΔefbΔecb mutant displayed reduced time-to-death and increased survival, as well as diminished abscess formation in organ tissues48. The ecb gene is found in all S. aureus genomes sequenced to date, whereas efb is found in many, but not all, human clinical isolates13.
SSLs also interfere with complement activation and phagocytosis. For example, SSL7 binds to human IgA and complement C5, interfering with IgA binding to FcαRI, the production of C5a and the oxidative burst of phagocytes in vitro; the in vivo contributions of SSL7 towards S. aureus pathogenesis are not known49. SSL10 binds to human and non-human primate IgG1, but not to immunoglobulins of lower vertebrates, and inhibits IgG1 binding to Fcγ receptors and the in vitro phagocytosis of IgG1-opsonized bacteria by immune cells50,51.
Staphylococcal binder of immunoglobulin (Sbi) is a secreted protein with two immunoglobulin binding domains (IgBDs; designated Sbi-I and Sbi-II), which are triple-helical bundles that associate with the Fcγ domain of human and vertebrate immunoglobulin (Box 2). Sbi-I and Sbi-II interfere with C1q binding to immunoglobulin and block the classical complement pathway52,53 (Fig. 1b). The Sbi-III and Sbi-IV domains associate with C3 and fH to form tripartite complexes that inhibit the alternative pathway54,55 (Fig. 1b). The sbi gene is located in the sbi–hlg locus of the core genome of all isolates13. Staphylokinase forms enzymatically active complexes with plasminogen, cleaving fibrin, defensins, human IgG, C3b and its proteolytically inactivated product iC3b on bacterial surfaces, thereby blocking complement activation56,57,58 (Fig. 1b). Collagen adhesin (Cna), a surface protein expressed by some S. aureus isolates, binds C1q and interferes with classical pathway activation, blocking the association between C1q and C1r59.
Neutrophil-mediated killing. Once phagocytosed, staphylococci are exposed to a variety of toxic products that kill and degrade the engulfed bacteria: antimicrobial peptides, nitric oxide (NO), ROS (that is, hydrogen peroxide, superoxide and hydroxyl radicals), cell wall hydrolases and proteolytic enzymes4. However, S. aureus has evolved a number of strategies to survive in this environment (Fig. 1c). Peptidoglycan acetylation (by the protein OatA), d-alanylation of teichoic acids (by the DltABCD complex), and lysyl- or alanyl-phosphatidylglycerol synthesis (by the protein MprF) provide staphylococcal resistance against lysozyme- and antimicrobial peptide-mediated killing by blocking enzymes (such as lysozyme) or peptides binding to the envelope target60,61,62. Staphyloxanthin, a carotenoid pigment synthesized by all S. aureus isolates63, provides resistance against hydrogen peroxide and/or hydroxyl radicals, the bactericidal compounds of neutrophils64 (this is not the case for CC75 isolates but we consider these to belong to a separate species, Staphylococcus argenteus (Box 1)). Similarly, two superoxide dismutases (SodA and SodM), fulfill overlapping functions in eliminating neutrophil superoxide65, whereas catalase (KatG) and alkylhydroperoxide reductase (AhpC) protect staphylococci against hydrogen peroxide66.
In response to nitrosative stress, S. aureus expresses flavohaemoglobin (Hmp), which detoxifies NO, and l-lactate dehydrogenase, which maintains redox-haemostasis and survival within neutrophils by producing l-lactate67. In addition to its role in inhibiting complement activation, Eap and two structural homologues, EapH1 and EapH2 (which do not inhibit complement), promote S. aureus survival by inhibiting neutrophil serine proteases (such as elastase, cathepsin G and proteinase 3)68. In the intravenous mouse challenge model, the S. aureus Δeap mutant displays a moderate virulence defect36; however, the ΔeapΔeapH1ΔeapH2 mutant displays reduced bacterial load and increased mouse survival68.
Staphylococcal killing of host cells. In addition to its ability to inhibit phagocyte-mediated killing, S. aureus also manipulates innate immune responses by inducing the killing of innate immune cells via PSMs and different toxins (Fig. 1d). The PSMα locus includes psma1–psma4, whereas PSMβ includes psmb1 and psmb2; psmb2 is found in only some, but not all, S. aureus strains13. Peptides similar to PSMα1–PSMα4 and PSMβ1 are expressed by Staphylococcus epidermidis, a commensal of the human skin that cannot cause abscess lesions or bloodstream infections in immune competent individuals25. Mutations that delete psma1–psma4 and psmb1 and psmb2 interfere with in vitro biofilm formation of S. aureus mutants and with the expression of virulence factors, including α-haemolysin25. S. aureus Δpsma1–psma4 mutants are attenuated in the mouse bloodstream infection model69, a phenotype that may be due to defects in biofilm formation, virulence gene expression and/or contributions of PSMα1–PSMα4 towards lysis of immune cells, presumably via membrane insertion and pore formation70.
β-barrel pore-forming toxins (β-PFTs) are secreted by the bacterium as soluble monomers and, on association with receptors on cell surfaces, assemble into multimeric pore structures, penetrating the lipid bilayer to invoke alterations in the physiology of injured cells or their outright lysis71. α-haemolysin (Hla), the prototype β-PFT of S. aureus, is encoded by the hla gene, which is located within IEC2. Although conserved among all S. aureus isolates, some lineages of S. aureus carry a nonsense mutation that blocks hla expression72. Hla binds to its receptor on host cells, ADAM10, and assembles into a heptameric pore; through the metalloproteinase activity of ADAM10, Hla modulates the function of immune cells, including neutrophils, or triggers lysis of epithelial cells73,74. S. aureus hla mutants display defects in disease severity in mouse models for lethal pneumonia, bacteraemia and SSTI; however, hla is not required for the establishment of S. aureus abscess lesions75,76,77. Based on ADAM10 expression on the surface of myeloid cells, organ epithelia and the vascular endothelium, Hla causes global, as well as organ-specific, changes to host physiology during S. aureus infection74.
Leukocidins are other β-PFTs secreted by S. aureus (Fig. 1d). Following leukocidin association with receptors on myeloid cells and erythrocytes, these toxins assemble from two different subunits (F and S) into an octameric pore structure78. All S. aureus strains produce at least three leukocidins, γ-haemolysin AB (HlgAB), HlgCB and leukocidin AB (LukAB; also known as LukGH), whereas other strains may also secrete Panton–Valentine leucocidin (PVL; which is encoded by lukPV) and LukED or LukMF79 (Box 1). The operon encoding LukAB is located immediately adjacent to hlb, whereas the operon encoding γ-haemolysin (hlgABC) is part of the sbi–hlg locus. LukAB binds to the I domain of human, but not mouse, αM integrin on myeloid cells80. Purified LukAB can trigger human neutrophils to release NETs that, at least temporarily, ensnare staphylococci81. LukAB has also been reported to promote S. aureus escape from the phagosome of neutrophils82. Purified HlgAB γ-haemolysin, but not purified HlgCB γ-haemolysin, is able to lyse human and rabbit red blood cells83. HlgAB binds to chemokine receptors CXCR1, CXCR2 and CC-chemokine receptor 2 (CCR2), whereas HlgCB uses complement receptors C5aR and C5L2 to associate with target cells84. Following staphylococcal inoculation into human blood, hlgABC is upregulated 34–145-fold85, and the S. aureus ΔhlgABC mutant displays reduced survival, presumably because HlgAB and HlgCB promote release of iron-compounds from erythrocytes, thereby enabling bacterial acquisition of this essential nutrient83. Both purified HlgAB and HlgCB promote lysis of neutrophils, monocytes and macrophages from humans, as well as non-human primates, and to a lesser degree rabbits and mice83. In a mouse intravenous challenge model, animals infected with a S. aureus ΔlukAB mutant displayed increased time-to-death and survival. Using subcutaneous inoculation in mice or rabbits, the S. aureus lukAB mutant did not display defects in skin abscess formation86. The ΔhlgAB mutant displayed a virulence defect in the intraperitoneal challenge model in mice84.
lukED is present in the GIβ locus of ∼70% of clinical S. aureus isolates13 (Box 1). Purified LukED triggers lysis of macrophages, dendritic cells and T cells from many different vertebrates, as the toxin binds to the chemokine receptors CCR5, CXCR1 and CXCR2 (Refs 87,88). For S. aureus strain Newman, which harbours GIβ, the ΔlukED mutation increased the time-to-death and survival of mice following intravenous challenge with mutant staphylococci89. PVL is secreted by S. aureus lysogenized with PVL phage90. PVL binds to the C5aR on neutrophils, monocytes and macrophages, but its activity is restricted towards human and rabbit cells9. By virtue of binding C5aR, PVL not only exerts its lytic activity on target host cells but can also facilitate the priming of human polymorphonuclear leukocytes by pro-inflammatory stimuli (for example, formyl peptides). Injection of purified recombinant PVL leads to increased immune cell recruitment and increased architectural destruction of the lung, owing to toxin-mediated recruitment and subsequent lysis of immune cells9. Only ∼2% of S. aureus isolates secrete PVL; however, community-acquired MRSA isolates frequently harbour PVL phages, and PVL expression is also associated with necrotizing pneumonia91. S. aureus ΔlukPV variants display defects in the pathogenesis of SSTIs and lung infections in rabbits, but not in mice, which seems to be due to neutrophil-mediated inflammatory responses and tissue distruction76,92. lukMF, genes for another phage-encoded leukocidin, are found in S. aureus isolates associated with bovine mastitis13.
Staphylococcal agglutination. Coagulation, the conversion of fibrinogen to a crosslinked fibrin meshwork by activated thrombin, is an innate defence of all vertebrates that immobilizes microbial invaders and attracts immune cells for phagocytic clearance of bacteria. Therefore, every successful bacterial pathogen must evolve mechanisms for escape from fibrin entrapment and subsequent phagocytosis by infiltrating immune cells. A hallmark of all S. aureus isolates is the secretion of two coagulases: coagulase (Coa) and von Willebrand factor-binding protein (vWbp)93. Coa and vWbp associate with prothrombin, a zymogen, to generate enzymatically active staphylothrombin, which cleaves the A and B peptides of fibrinogen to generate fibrin fibrils94 (Fig. 2). As staphylothrombin does not cleave other substrates of thrombin, it avoids the activation of clotting and inflammatory factors that ordinarily accompany fibrin polymerization95. Staphylothrombin activity is not subject to feedback inhibition through host antithrombin. However, staphylothrombin is blocked by dabigatran and other direct thrombin inhibitors of the same family96. The staphylothrombin-generated fibrin meshwork protects S. aureus from phagocytes and contributes to the formation of staphylococcal abscess lesions and lethal bacteraemia in mice97. Activation of prothrombin is mediated by the D1 and D2 domains in the N-terminal region of Coa and is blocked by specific antibodies, which provide protection from S. aureus bloodstream infection in the mouse model98. Perhaps owing to purifying selection, coa is one of the most variable genes in the core genome of S. aureus, with >50% sequence variation in the coding sequence for its D1–D2 domains and 14 distinct isoforms93 (Table 1). vWbp also has conserved D1–D2 domains for association with prothrombin, but this complex generates fibrin at a reduced rate and contributes to abscess formation without affecting staphylococcal escape from phagocytosis99. The gene encoding vWbp, vwb, has limited sequence variability98.
Physiological host defences immobilize bacteria through the activation of the serine protease zymogens prothrombin (also known as factor II) and factor X (not shown). In the contact-activation pathway, surface contact results in the autocleavage of prothrombin, thereby generating thrombin (also known as factor IIa). Staphylococcus aureus superantigen-like 10 (SSL10) inhibits prothrombin autoactivation, whereas the S. aureus coagulases, coagulase (Coa) and von Willebrand factor-binding protein (vWbp) convert prothrombin to staphylothrombin. Both thrombin and staphylothrombin cleave fibrinopeptides A and B from fibrinogen to generate fibrin, which self-assembles and polymerizes into cable structures that immobilize bacteria. Thrombin activation results in the activation of additional haemostasis factors that facilitate the simultaneous attraction of phagocytes to immobilized bacteria, which is thus inhibited by SSL10 secretion. However, staphylothrombin cleaves fibrinopeptides from fibrinogen without activation of other haemostasis factors and promotes fibrin polymer assembly on the staphylococcal surface, where it protects the bacterium from neutrophils and phagocytic clearance. Fibrin agglutination on the staphylococcal surface also involves the S. aureus surface proteins clumping factor A (ClfA), fibronectin-binding protein A (FnBPA) and FnBPB, which bind to the fibrinogen γ-chains.
S. aureus agglutinates with Coa- or vWbp-derived fibrin fibrils, which requires clumping factor A (ClfA), a glycosylated, sortase-anchored surface protein, the immunoglobulin-like domains of which bind to the carboxy-terminal end of the γ-chain in fibrinogen and fibrin (that is, the D domain)96,100,101 (Fig. 2). Thus, ClfA acts synergistically with Coa or vWbp in protecting staphylococci from opsonophagocytic killing, and ΔclfA mutants display defects in the pathogenesis of lethal bloodstream infections in mice96.
Four other sortase-anchored surface proteins use their immunoglobulin-like domains to bind fibrinogen or fibrin: ClfB (which binds to the fibrinogen α-chain), fibronectin-binding protein A (FnBPA) and FnBPB (which bind to the C-terminal end of the fibrinogen γ-chain), and bone sialoprotein binding protein (Bbp; also known as SdrE isoform; which binds to the fibrinogen α-chain)102 (Fig. 2). These surface proteins display functional redundancy for the S. aureus agglutination pathway and contribute to the pathogenesis of bloodstream infections103.
Purified, recombinant SSL10 has also been reported to bind human fibrinogen and fibronectin, as well as pig prothrombin and factor Xa51. The association of SSL10 with prothrombin and factor Xa occurs via the γ-carboxylic acid (Gla) domain and interferes with calcium-activated blood clotting but not with staphylothrombin-mediated fibrin formation51 (Fig. 2). ssl7 and ssl10 are not found in all S. aureus isolates13.
Staphylokinase activates human, but not mouse, plasminogen and may solubilize coagulase-induced fibrin deposits, thereby aiding S. aureus in generating purulent lesions for dissemination to new hosts (Fig. 1b). Of note, binding of surface proteins by fibrinogen or fibrin deposits has also been reported to influence staphylococcal interference with platelet aggregation and innate immune functions104,105. In addition to their role in binding to fibrin and fibrinogen, some surface proteins have been reported to bind additional host ligands, at least in vitro. Binding to these additional host ligands, which include complement factor I (by the surface protein ClfA), fibronectin (by the surface proteins FnBPA and FnBPB), keratin 10 (by the surface protein ClfB) and loricrin (by the surface protein ClfB), may contribute to staphylococcal immune evasion, invasion of host cells or colonization of squamous epithelia102. fnbpA and fnbpB, which are components of the S. aureus core genome, have sequence polymorphisms, and seven isotypes with discrete antigenicity have been described106 (Box 1).
Adenosine and deoxyadenosine signalling and NETosis. Adenosine is a potent mediator of immune responses and, under physiological conditions, is synthesized following hypoxia, exposure to ROS and cell lysis associated with tissue damage. Adenosine elicits its biological effects by binding to one or more of four G protein-coupled receptors107: A1, A2A, A2B and A3. Adenosine receptor interaction triggers anti-inflammatory signalling cascades that inhibit platelet aggregation, neutrophil superoxide burst, neutrophil degranulation, T cell activation and release of the cytokines IL-1α and IL-10 (Ref. 108). S. aureus increases the concentrations of extracellular adenosine during infection by expressing adenosine synthase A (AdsA), a sortase-anchored protein that catalyses the dephosphorylation of adenosine mono-, di- and triphosphates109 (Fig. 3a). Both ex vivo and during mouse infection, the ΔadsA mutation increases killing of staphylococci by blood neutrophils, while decreasing extracellular adenosine109. Thus, AdsA-mediated synthesis of adenosine promotes survival of S. aureus within neutrophils, presumably by inhibiting superoxide burst and/or degranulation109. Furthermore, adenosine decreases major histocompatibility complex (MHC) class II expression by macrophages and dendritic cells and dampens IL-12 production, a pivotal stimulus for T helper 1-type immune responses107. Staphylococcal enhancement of adenosine production may therefore interfere with T cell effector mechanisms and adaptive immune responses in infected hosts109.
a | Staphylococcus aureus infection and its associated inflammatory damage promote the release of ATP, which is converted by adenosine synthase A (AdsA) into the immune suppressive signalling molecule adenosine (A). Adenosine inhibits activation of B cells, T cells, macrophages and dendritic cells via adenosine receptor (AdoR) signalling by acting on four different receptors (AdoR1, AdoR2A, AdoR2B and AdoR3). Under physiological conditions, CD39 and CD73 generate adenosine signals to limit inflammatory responses; CD39 and CD73 are also responsible for the adenosine halo surrounding immune cells and for immune suppressive states involving regulatory T cells (T cells expressing the FOXP3+ marker protein (not shown)). b | S. aureus induced NETosis of infiltrating neutrophils leads to nuclease-mediated degradation of the DNA fibres that are the major components of neutrophil extracellular traps (NETs) and AdsA-mediated conversion of 5′-monophosphate-deoxyadenosine (dAMP) into deoxyadenosine (dAdo), which promotes autocleavage of the apoptosis factor pro-capsase 3 to caspase 3. Caspase 3 induces macrophage death, thereby protecting S. aureus against professional phagocytes. IL-12, interleukin-12; MHC II, major histocompatibility complex class II; Nuc, staphylococcal nuclease; PMN, polymorphonuclear leukocyte.
AdsA activity also modulates immune responses following the degradation of NETs. During bloodstream infection in mice, S. aureus disseminates to many different organ tissues to establish abscess lesions. These lesions are composed of a bacterial nidus, designated as the staphylococcal abscess community (SAC), encased within a pseudocapsule of fibrin deposits, and surrounded by layers of immune cells97. In spite of large numbers of infiltrated neutrophils, mice are unable to eliminate staphylococci from abscess lesions and eventually succumb to persistent infection36. Although neutrophils use NETosis to entangle staphylococci, NETs are degraded by staphylococcal nuclease (Nuc) and thereby fail to exert bactericidal activities110 (Fig. 3b). Nucdigestion of NETs releases 5′ and 3′ monophosphate nucleotides that are converted by AdsA into deoxyadenosine111 (Fig. 3b). Deoxyadenosine production triggers caspase 3 induced apoptosis of macrophages and prevents phagocyte entry into the SAC, the core of staphylococcal abscess lesions, thereby promoting bacterial survival within the lesion111.
Manipulation of adaptive immune responses
B cell responses. S. aureus is capable of manipulating B cell survival and function, especially through production of staphylococcal protein A (SpA), which is a sortase-anchored surface protein with high affinity for vertebrate immunoglobulin, including human IgA, IgD, IgG1–IgG4, IgM and IgE112. SpA is initially deposited in the staphylococcal envelope and subsequently released by cell wall hydrolases (LytM)113. spa is expressed by all clinical S. aureus isolates; the immunoglobulin binding domains are conserved in the genomes of these isolates, but region X, the cell wall spanning domain of SpA, is a highly polymorphic sequence114,115 (Box 1).
The immunosuppressive attributes of SpA have been ascribed to two distinct binding activities: association with the Fcγ domain and with the Fab domains of antibodies116,117. SpA binding to the Fcγ domain of IgG blocks phagocytosis of staphylococci118, whereas SpA binding to the Fab domains and crosslinking of VH3 clan IgM promotes B cell superantigen activity119 (Fig. 4a). Of note, SpA binds specifically to VH3 clan IgM antibodies, which mediate the predominant antibody responses to infection and immunization, but not to other clan antibodies. In the intravenous challenge model of S. aureus infected mice, spa expression suppresses antibody responses against many different staphylococcal antigens and provides antiphagocytic attributes, promoting staphylococcal survival in blood120. Infection of mice with S. aureus spa variants that cannot bind immunoglobulin is associated with attenuated disease and with antibody responses against many different antigens that can protect animals against subsequent lethal challenge with other S. aureus isolates120.
a | Staphylococcus aureus releases staphylococcal protein A (SpA) into host tissues, where it binds to and crosslinks VH3 clan B cell receptors. In B1 cells, marginal zone (MZ) B cells and B2 cells, SpA crosslinking is associated with proliferative expansion and apoptotic collapse. The death of these cells impedes the development of adaptive immunity during S. aureus infections. b | In VH3+ plasmablasts, SpA crosslinking promotes somatic hypermutation and class switching from IgM antibodies to IgG antibodies, followed by the secretion of antibodies that are not specific for the S. aureus antigen. c | S. aureus secretes T cell superantigen (SAg), which crosslinks major histocompatibility complex class II antigens (MHC II) on the surface of antigen-presenting cells and T cell receptors (TCRs) on the surface of T helper (TH) cells, triggering T cell expansion and anergy and causing cytokine storms (including interleukin-2 (IL-2), interferon-γ (IFNγ), IL-1β and tumour necrosis factor (TNF)). As a result, a T cell response specific for S. aureus antigens is not produced.
Mice harbour a limited repertoire of VH3+ B cells, whereas humans possess large populations of these cells, yet both species cannot develop SpA-neutralizing antibodies during infection121. S. aureus infection in humans triggers expansions of VH3 idiotypic plasmablasts (>90% of blood plasmablasts), the antibodies of which (that is, the B cell receptors) associate with SpA via their Fab domains but do not display pathogen-specific binding activities121 (Fig. 4b). When mice are treated with purified SpA, crosslinking of VH3 clonal B cells triggers proliferation and apoptotic collapse of expanded populations of B cells122. It is not clear, however, whether apoptotic collapse of expanded lymphocyte populations occurs during S. aureus infection in mice or in humans.
Non-toxigenic SpA, designated SpAKKAA, was engineered by substituting twenty amino acid residues essential for its association with the Fcγ and Fab regions123. Although SpAKKAA has twenty amino acid substitutions, this antigen elicits antibodies that neutralize SpA when injected into animals123. The SpAKKAA-derived polyclonal antibodies promote phagocytosis of staphylococci and display adjuvant attributes by suppressing staphylococcal B cell superantigen activity and promoting humoral immune responses against a wide range of S. aureus antigens123. Studies with mouse monoclonal antibodies corroborate this concept53.
T cell responses. Staphylococcal T cell superantigens bind to MHC class II molecules on the surface of antigen-presenting cells, providing antigen-independent crosslinking with T cell receptors on T helper cells124 (Fig. 4c). S. aureus strains have been shown to express 23 different enterotoxins and T cell superantigens125. Three superantigens are most frequently associated with human disease — toxic shock syndrome toxin 1 (TSST1), staphylococcal enterotoxin B (SEB) and SEC — each providing high-affinity interactions with distinct subsets of Vβ chain T cell receptors126. In humans with toxic shock syndrome, S. aureus secretion of TSST1 or other enterotoxins trigger expansions of cognate T cell populations, up to 30% of blood lymphocytes and nonspecific release of cytokines, preventing a focused adaptive immune response127. Depending on the site and severity of S. aureus infection or intoxication, superantigen-mediated activation of T cell responses may be associated with cytokine storms and toxic shock syndrome pathology128. Staphylococcal superantigens are also thought to interfere with antigen-specific proliferation of T cells and with antibody responses against specific subsets of staphylococcal antigens, including staphylococcal superantigens129. It is not yet known whether superantigens have a crucial role in the suppression of T cell responses in mice that are observed during S. aureus bloodstream infections130.
S. aureus can also manipulate T cell responses by promoting T cell lysis. For example, δ-toxin (Hld; also known as δ-haemolysin), a member of the PSMα family, can lyse T cells131 and has also been reported to trigger mast cell degranulation, which could be a key factor in the exacerbation of S. aureus infected atopic dermatitis lesions, where histamine release is otherwise triggered by antigen-induced crosslinking of IgE bound to FcγRI receptor132. Hld is encrypted within the agr-regulated RNA III molecule, the regulatory arm of staphylococcal quorum-sensing133.
Outlook
S. aureus strains secrete many immune evasive molecules and, when placed under selective pressure, acquire mobile genetic elements with additional factors so that staphylococci meet the demands for invasion of host species and replication in specific anatomical niches13. These strategies are accomplished through factors that block phagocyte chemotaxis, complement activation, phagocytic uptake and oxidative killing, often redirecting host defences such as fibrin formation or NETosis to favour pathogen replication. Staphylococcal infection is also associated with perturbations of adaptive responses, including the disruptive proliferation of B cells and T cells, which prevents the establishment of protective immune responses. Why do S. aureus isolates acquire so many different immune evasion factors when other bacterial pathogens manage with only a small number? Addressing this question, one should consider that S. aureus maintains life-long associations with its human hosts, colonizing and reiteratively invading large segments of the population. We know of no other bacterial pathogen able to sustain a similar lifestyle. Staphylococcal capabilities of causing recurrent and reiterative infections probably rely on two mechanisms. First, S. aureus manipulation of B cell and T cell responses must be successful, as increased age is not associated with decreased incidence. Second, mobile genetic elements enable acquisition and/or exchange of immune evasive traits between S. aureus strains and horizontal gene transfer may implement disease in individuals who did mount successful immune responses against immune evasion determinants. If pressed to predict the future of S. aureus as it evolves with a population whose lifespan continues to increase, we would forecast more invasion by mobile genetic elements and more immune evasion determinants.
Considering the formidable weapons of the pathogen against the host's immune defences, development of vaccines against S. aureus is a daunting task. Conventional approaches for vaccine development follow the mantra of eliciting specific antibodies that trigger pathogen killing in vitro and disease protection in preclinical (animal) models of staphylococcal disease. The simplest means of achieving these goals are whole-cell vaccines, either killed or attenuated preparations. Indeed, use of an autologous vaccine for individuals with recurrent S. aureus infection has been practised for many years134. This approach can elicit pathogen-specific antibodies; however, it has not been demonstrated to raise protective immunity135. Live-attenuated vaccines have been studied in animal models with variable success but not in humans. If one considers that the immune-evasive strategies are often species specific, it seems risky to derive claims on human protective immune responses against S. aureus from experiments with animals. What is true for whole-cell vaccines certainly applies to subunit vaccines. Antibodies against capsular polysaccharide, ClfA, IsdB and lipoteichoic acid bind to the surface of the pathogen, which enables phagocyte-mediated killing in vitro and provides protection from infection in specific animal models; however, the corresponding vaccines or antibodies did not achieve efficacy in clinical trials136,137,138. It occurred to us that in vitro assays for phagocytic killing of S. aureus often do not consider key evasion strategies of the pathogen and the corresponding defences of humans. We believe this can be addressed with Lancefield's assay for antibody-mediated killing of bacteria in fresh anti-coagulated human blood139. The Lancefield assay can also be used for prospective studies in humans, assessing antibody titres, status of immunity (bactericidal activity of blood) and probability of disease. This approach may identify criteria for protective immunity, stratify patients at risk for S. aureus disease and guide applications for immune-therapy or vaccination that reduce the incidence of disease. Previous work targeted B cell and T cell superantigens from S. aureus as vaccine antigens to enable the development of broad-spectrum immune responses during each encounter of the host with this pathogen123,140. Another promising approach exploited the structural relatedness of immune evasion factors to target multiple molecules with antibodies that recognize conserved structural features141 (Box 2). Nonetheless, the efficacy of these approaches has not yet been assessed in human clinical trials.
References
van Belkum, A. et al. Co-evolutionary aspects of human colonisation and infection by Staphylococcus aureus. Infect. Genet. Evol. 9, 32–47 (2009).
David, M. Z. & Daum, R. S. Community-associated methicillin-resistant Staphylococcus aureus: epidemiology and clinical consequences of an emerging epidemic. Clin. Microbiol. Rev. 23, 616–687 (2010).
Kallen, A. J. et al. Health care-associated invasive MRSA infections, 2005–2008. JAMA 304, 641–648 (2010).
Spaan, A. N., Surewaard, B. G. J., Nijland, R. & van Strijp, J. A. G. Neutrophils versus Staphylococcus aureus: a biological tug of war. Annu. Rev. Microbiol. 67, 629–650 (2013). This excellent review summarizes the molecular events that occur during encounters between neutrophils and staphylococci.
Curnutte, J. T., Whitten, D. M. & Babior, B. M. Defective superoxide production by granulocytes from patients with chronic granulomatous disease. N. Engl. J. Med. 290, 593–597 (1974).
Casanova, J. L., Holland, S. M. & Notarangelo, L. D. Inborn errors of human JAKs and STATs. Immunity 36, 515–528 (2012).
Maródi, L. et al. Molecular mechanisms of mucocutaneous immunity against Candida and Staphylococcus species. J. Allergy Clin. Immunol. 130, 1019–1027 (2012).
Chemaly, R. F. et al. Characteristics and outcomes of methicillin-resistant Staphylococcus aureus surgical-site infections in patients with cancer: a case-control study. Ann. Surg. Oncol. 17, 1499–1506 (2010).
Spaan, A. N. et al. The staphylococcal toxin panton-valentine leukocidin targets human c5a receptors. Cell Host Microbe 13, 584–594 (2013). This report describes the C5a receptor as the target of PVL, an important leukocidin secreted by some S. aureus strains.
Williams, R. J. et al. Identification of a novel gene cluster encoding staphylococcal exotoxin-like proteins: characterization of the prototypic gene and its protein product, SET1. Infect. Immun. 68, 4407–4415 (2000).
Fitzgerald, J. R. et al. Genome diversification in Staphylococcus aureus: molecular evolution of a highly variable chromosomal region encoding the staphylococcal exotoxin-like family of proteins. Infect. Immun. 71, 2827–2838 (2003).
Arcus, V. L., Langley, R., Proft, T., Fraser, J. D. & Baker, E. N. The three-dimensional structure of a superantigen-like protein, SET3, from a pathogenicity island of the Staphylococcus aureus genome. J. Biol. Chem. 277, 32274–32281 (2002).
McCarthy, A. J. & Lindsay, J. A. Staphylococcus aureus innate immune evasion is lineage-specific: a bioinformatics study. Infect. Genet. Evol. 19, 7–14 (2013). This report characterizes lineage-specific alleles and distribution of S. aureus immune evasion determinants.
Bestebroer, J. et al. Staphylococcal superantigen-like 5 binds PSGL-1 and inhibits P-selectin-mediated neutrophil rolling. Blood 109, 2936–2943 (2007).
Chung, M. C. et al. The crystal structure of staphylococcal superantigen-like protein 11 in complex with sialyl Lewis X reveals the mechanism for cell binding and immune inhibition. Mol. Microbiol. 66, 1342–1355 (2007).
Bestebroer, J. et al. Staphylococcal SSL5 inhibits leukocyte activation by chemokines and anaphylatoxins. Blood 113, 328–337 (2009).
de Haas, C. J. et al. Staphylococcal superantigen-like 5 activates platelets and supports platelet adhesion under flow conditions, which involves glycoprotein Ibα and αIIbβ3. J. Thromb. Haemost. 7, 1867–1874 (2009).
Hu, H. et al. GPVI and GPIbα mediate staphylococcal superantigen-like protein 5 (SSL5) induced platelet activation and direct toward glycans as potential inhibitors. PLoS ONE 6, e19190 (2011).
Armstrong, P. C. et al. Staphylococcal superantigen-like protein 5 induces thrombotic and bleeding complications in vivo: inhibition by an anti-SSL5 antibody and the glycan bimosiamose. J. Thromb. Haemost. 10, 2607–2609 (2012).
Itoh, S. et al. Staphylococcal superantigen-like protein 5 inhibits matrix metalloproteinase 9 from human neutrophils. Infect. Immun. 78, 3298–3305 (2010).
Hermans, S. J. et al. Structural and functional properties of staphylococcal superantigen-like protein 4. Infect. Immun. 80, 4004–4013 (2012).
Yokoyama, R. et al. Staphylococcal superantigen-like protein 3 binds to the Toll-like receptor 2 extracellular domain and inhibits cytokine production induced by Staphylococcus aureus, cell wall component, or lipopeptides in murine macrophages. Infect. Immun. 80, 2816–2825 (2012).
Fevre, C. et al. Staphylococcus aureus proteins SSL6 and SElX interact with neutrophil receptors as identified using secretome phage display. Cell. Microbiol. 16, 1646–1665 (2014).
Chavakis, T. et al. Staphylococcus aureus extracellular adherence protein serves as anti-inflammatory factor by inhibiting the recruitment of host leukocytes. Nat. Med. 8, 687–693 (2002).
Peschel, A. & Otto, M. Phenol-soluble modulins and staphylococcal infection. Nat. Rev. Microbiol. 11, 667–673 (2013).
Kretschmer, D. et al. Human formyl peptide receptor 2 senses highly pathogenic Staphylococcus aureus. Cell Host Microbe 7, 463–473 (2010). This paper identifies formyl receptor 2 signalling as the target for S. aureus phenol-soluble modulins.
Forsman, H., Christenson, K., Bylund, J. & Dahlgren, C. Receptor-dependent and -independent immunomodulatory effects of phenol-soluble modulin peptides from Staphylococcus aureus on human neutrophils are abrogated through peptide inactivation by reactive oxygen species. Infect. Immun. 80, 1987–1995 (2012).
Cheung, G. Y. et al. Production of an attenuated phenol-soluble modulin variant unique to the MRSA clonal complex 30 increases severity of bloodstream infection. PLoS Pathog. 10, e1004298 (2014).
van Wamel, W. J., Rooijakkers, S. H., Ruyken, M., van Kessel, K. P. & van Strijp, J. A. The innate immune modulators staphylococcal complement inhibitor and chemotaxis inhibitory protein of Staphylococcus aureus are located on β-hemolysin-converting bacteriophages. J. Bacteriol. 188, 1310–1315 (2006).
de Haas, C. J. et al. Chemotaxis inhibitory protein of Staphylococcus aureus, a bacterial antiinflammatory agent. J. Exp. Med. 199, 687–695 (2004). This report characterizes CHIPS as an inhbitor of chemotaxis and complement-mediated immune cell activation.
Postma, B. et al. Chemotaxis inhibitory protein of Staphylococcus aureus binds specifically to the C5a and formylated peptide receptor. J. Immunol. 172, 6994–7001 (2004).
Prat, C., Bestebroer, J., de Haas, C. J., van Strijp, J. A. & van Kessel, K. P. A new staphylococcal anti-inflammatory protein that antagonizes the formyl peptide receptor-like 1. J. Immunol. 177, 8017–8026 (2006).
Prat, C. et al. A homolog of formyl peptide receptor-like 1 (FPRL1) inhibitor from Staphylococcus aureus (FPRL1 inhibitory protein) that inhibits FPRL1 and FPR. J. Immunol. 183, 6569–6578 (2009).
Laarman, A. J. et al. Staphylococcus aureus staphopain A inhibits CXCR2-dependent neutrophil activation and chemotaxis. EMBO J. 31, 3607–3619 (2012).
Bubeck Wardenburg, J., Williams, W. A. & Missiakas, D. Host defenses against Staphylococcus aureus infection require recognition of bacterial lipoproteins. Proc. Natl Acad. Sci. USA 103, 13831–13836 (2006).
Cheng, A. G. et al. Genetic requirements for Staphylococcus aureus abscess formation and persistence in host tissues. FASEB J. 23, 3393–3404 (2009).
Gros, P., Milder, F. J. & Janssen, B. J. Complement driven by conformational changes. Nat. Rev. Immunol. 8, 48–58 (2008).
Laarman, A., Milder, F., van Strijp, J. A. & Rooijakkers, S. H. Complement inhibition by gram-positive pathogens: molecular mechanisms and therapeutic implications. J. Mol. Med. 88, 115–120 (2010).
O'Riordan, K. & Lee, J. C. Staphylococcus aureus capsular polysaccharides. Clin. Microbiol. Rev. 17, 218–234 (2004).
Montgomery, C. P. et al. Comparison of virulence in community-associated methicillin-resistant Staphylococcus aureus pulsotypes USA300 and USA400 in a rat model of pneumonia. J. Infect. Dis. 198, 561–570 (2008).
Laarman, A. J. et al. Staphylococcus aureus metalloprotease aureolysin cleaves complement C3 to mediate immune evasion. J. Immunol. 186, 6445–6453 (2011).
Rooijakkers, S. H. et al. Immune evasion by a staphylococcal complement inhibitor that acts on C3 convertases. Nat. Immunol. 6, 920–927 (2005). The paper discovers SCIN as an inhibitor of C3 convertases, blocking C3b deposition on staphylococcal surfaces.
Jongerius, I. et al. Staphylococcal complement evasion by various convertase-blocking molecules. J. Exp. Med. 204, 2461–2471 (2007). This manuscript identifies SCIN-B, SCIN-C, Ecb and Efb as inhibitors of C3 and C5 convertases.
Chen, H. et al. Allosteric inhibition of complement function by a staphylococcal immune evasion protein. Proc. Natl Acad. Sci. USA 107, 17621–17626 (2010).
Amdahl, H. et al. Staphylococcal Ecb protein and host complement regulator factor H enhance functions of each other in bacterial immune evasion. J. Immunol. 191, 1775–1184 (2013).
Jongerius, I., Garcia, B. L., Geisbrecht, B. V., van Strijp, J. A. & Rooijakkers, S. H. Convertase inhibitory properties of staphylococcal extracellular complement-binding protein. J. Biol. Chem. 285, 14973–14979 (2010).
Ko, Y. P. et al. Phagocytosis escape by a Staphylococcus aureus protein that connects complement and coagulation proteins at the bacterial surface. PLoS Pathog. 9, e1003816 (2013).
Jongerius, I. et al. Staphylococcus aureus virulence is enhanced by secreted factors that block innate immune defenses. J. Innate Immun. 4, 301–311 (2012).
Bestebroer, J. et al. Functional basis for complement evasion by staphylococcal superantigen-like 7. Cell. Microbiol. 12, 1506–1516 (2010).
Patel, D., Wines, B. D., Langley, R. J. & Fraser, J. D. Specificity of staphylococcal superantigen-like protein 10 toward human IgG1 Fc domain. J. Immunol. 184, 6283–6292 (2010).
Itoh, S. et al. Staphylococcal superantigen-like protein 10 (SSL10) inhibits blood coagulation by binding to prothrombin and factor Xa via their γ-carboxyglutamic acid (Gla) domain. J. Biol. Chem. 288, 21569–21580 (2013).
Zhang, L., Jacobsson, K., Vasi, J., Lindberg, M. & Frykberg, L. A second IgG-binding protein in Staphylococcus aureus. Microbiology 144, 985–991 (1998).
Kim, H. K. et al. Protein A-specific monoclonal antibodies and the prevention of Staphylococcus aureus disease in mice. Infect. Immun. 80, 3460–3470 (2012).
Burman, J. D. et al. Interaction of human complement with Sbi, a staphylococcal immunoglobulin-binding protein: indications of a novel mechanism of complement evasion by Staphylococcus aureus. J. Biol. Chem. 283, 17579–17593 (2008).
Haupt, K. et al. The Staphylococcus aureus protein Sbi acts as a complement inhibitor and forms a tripartite complex with host complement Factor H and C3b. PloS Pathog. 4, e1000250 (2008). The manuscript explains the functional properties of Sbi-mediated immune evasion.
Parry, M. A. et al. The ternary microplasmin-staphylokinase-microplasmin complex is a proteinase-cofactor-substrate complex in action. Nat. Struct. Biol. 5, 917–923 (1998).
Rooijakkers, S. H., van Wamel, W. J., Ruyken, M., van Kessel, K. P. & van Strijp, J. A. Anti-opsonic properties of staphylokinase. Microbes Infect. 7, 476–484 (2005).
Jin, T. et al. Staphylococcus aureus resists human defensins by production of staphylokinase, a novel bacterial evasion mechanism. J. Immunol. 172, 1169–1176 (2004).
Kang, M. et al. Collagen-binding microbial surface components recognizing adhesive matrix molecule (MSCRAMM) of Gram-positive bacteria inhibit complement activation via the classical pathway. J. Biol. Chem. 288, 20520–20531 (2013).
Bera, A., Herbert, S., Jakob, A., Vollmer, W. & Götz, F. Why are pathogenic staphylococci so lysozyme resistant? The peptidoglycan O-acetyltransferase OatA is the major determinant for lysozyme resistance of Staphylococcus aureus. Mol. Microbiol. 55, 778–787 (2005).
Peschel, A. et al. Inactivation of the dlt operon in Staphylococcus aureus confers sensitivity to defensins, protegrins, and other antimicrobial peptides. J. Biol. Chem. 274, 8405–8410 (1999).
Ernst, C. M. & Peschel, A. Broad-spectrum antimicrobial peptide resistance by MprF-mediated aminoacylation and flipping of phospholipids. Mol. Microbiol. 80, 290–299 (2011).
Pelz, A. et al. Structure and biosynthesis of staphyloxanthin from Staphylococcus aureus. J. Biol. Chem. 280, 32493–32498 (2005).
Liu, G. Y. et al. Staphylococcus aureus golden pigment impairs neutrophil killing and promotes virulence through its antioxidant activity. J. Exp. Med. 202, 209–215 (2005).
Karavolos, M. H., Horsburgh, M. J., Ingham, E. & Foster, S. J. Role and regulation of the superoxide dismutases of Staphylococcus aureus. Microbiology 149, 2749–2758 (2003).
Cosgrove, K. et al. Catalase (KatA) and alkyl hydroperoxide reductase (AhpC) have compensatory roles in peroxide stress resistance and are required for survival, persistence, and nasal colonization in Staphylococcus aureus. J. Bacteriol. 189, 1025–1035 (2007).
Richardson, A. R., Libby, S. J. & Fang, F. C. A nitric oxide-inducible lactate dehydrogenase enables Staphylococcus aureus to resist innate immunity. Science 319, 1672–1676 (2008). A report on how staphylococcal metabolism diverts nitrosative stress and provides escape from phagocytic killing.
Stapels, D. A. et al. Staphylococcus aureus secretes a unique class of neutrophil serine protease inhibitors. Proc. Natl Acad. Sci. USA 111, 13187–13192 (2014). This paper shows that three secreted molecules of S. aureus — Eap, EapH1 and EapH2 — block neutrophil proteases to escape phagocytic killing.
Wang, R. et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 13, 1418–1420 (2007).
Surewaard, B. G. et al. Staphylococcal α-phenol soluble modulins contribute to neutrophil lysis after phagocytosis. Cell. Microbiol. 15, 1427–1437 (2013).
Heuck, A. P., Tweten, R. K. & Johnson, A. E. β-barrel pore-forming toxins: intriguing dimorphic proteins. Biochemistry 40, 9065–9073 (2001).
DeLeo, F. R. et al. Molecular differentiation of historic phage-type 80/81 and contemporary epidemic Staphylococcus aureus. Proc. Natl Acad. Sci. USA 108, 18091–18096 (2011).
Wilke, G. A. & Bubeck Wardenburg, J. Role of a disintegrin and metalloprotease 10 in Staphylococcus aureus α-hemolysin-mediated cellular injury. Proc. Natl Acad. Sci. USA 107, 13473–13478 (2010). The paper identifies ADAM10 as the receptor for staphylococcal α -haemolysin.
Becker, R. E., Berube, B. J., Sampedro, G. R., Dedent, A. C. & Bubeck Wardenburg, J. Tissue-specific patterning of host innate immune responses by Staphylococcus aureus α-toxin. J. Innate Immun. 6, 619–631 (2014).
Bubeck Wardenburg, J. & Schneewind, O. Vaccine protection against Staphylococcus aureus pneumonia. J. Exp. Med. 205, 287–294 (2008).
Bubeck Wardenburg, J., Palazzolo-Ballance, A. M., Otto, M., Schneewind, O. & DeLeo, F. R. Panton-Valentine leukocidin is not a virulence determinant in murine models of community-associated methicillin-resistant Staphylococcus aureus disease. J. Infect. Dis. 198, 1166–1170 (2008).
Powers, M. E., Kim, H. K., Wang, Y.-T. & Bubeck-Wardenburg, J. ADAM10 mediates vascular injury induced by Staphylococcus aureus α-hemolysin. J. Infect. Dis. 206, 352–356 (2012).
Miles, G., Movileanu, L. & Bayley, H. Subunit composition of a bicomponent toxin: staphylococcal leukocidin forms an octameric transmembrane pore. Protein Sci. 11, 894–902 (2002).
Alonzo, F. & Torres, V. J. The bicomponent pore-forming leucocidins of Staphylococcus aureus. Microbiol. Mol. Biol. Rev. 78, 199–230 (2014).
DuMont, A. L. et al. Staphylococcus aureus LukAB cytotoxin kills human neutrophils by targeting the CD11b subunit of the integrin Mac-1. Proc. Natl Acad. Sci. USA 110, 10794–10799 (2013).
Malachowa, N., Kobayashi, S. D., Freedman, B., Dorward, D. W. & DeLeo, F. R. Staphylococcus aureus leukotoxin GH promotes formation of neutrophil extracellular traps. J. Immunol. 191, 6022–6029 (2013).
DuMont, A. L. et al. Staphylococcus aureus elaborates leukocidin AB to mediate escape from within human neutrophils. Infect. Immun. 81, 1830–1841 (2013).
Malachowa, N. & DeLeo, F. R. Staphylococcus aureus survival in human blood. Virulence 2, 567–569 (2011). This manuscript identifies the receptors for staphylococcal γ -haemolysins AB and CB on immune cells.
Spaan, A. N. et al. The staphylococcal toxins γ-haemolysin AB and CB differentially target phagocytes by employing specific chemokine receptors. Nat. Commun. 5, 5438 (2014).
Malachowa, N. et al. Global changes in Staphylococcus aureus gene expression in human blood. PLoS ONE 6, e18617 (2011).
Malachowa, N. et al. Staphylococcus aureus leukotoxin GH promotes inflammation. J. Infect. Dis. 206, 1185–1193 (2012).
Alonzo, F. et al. CCR5 is a receptor for Staphylococcus aureus leukotoxin ED. Nature 493, 51–55 (2013). This paper implicated G protein-coupled receptors as the targets of leukocidins, which are β -PFTs that are secereted by S. aureus to subvert innate immune responses.
Reyes-Robles, T. et al. Staphylococcus aureus leukotoxin ED targets the chemokine receptors CXCR1 and CXCR2 to kill leukocytes and promote infection. Cell Host Microbe 14, 453–459 (2013).
Alonzo, F. et al. Staphylococcus aureus leucocidin ED contributes to systemic infection by targeting neutrophils and promoting bacterial growth in vivo. Mol. Microbiol. 83, 423–435 (2012).
Panton, P. N. & Valentine, F. C. O. Staphylococcal toxin. Lancet 222, 506–508 (1932).
Gillet, Y. et al. Association between Staphylococcus aureus strains carrying gene for Panton–Valentine leukocidin and highly lethal necrotising pneumonia in young immunocompetent patients. Lancet 359, 753–759 (2002).
Diep, B. A. et al. Polymorphonuclear leukocytes mediate Staphylococcus aureus Panton–Valentine leukocidin-induced lung inflammation and injury. Proc. Natl Acad. Sci. USA 107, 5587–5592 (2010).
McCarthy, A. J. & Lindsay, J. A. Genetic variation in Staphylococcus aureus surface and immune evasion genes is lineage associated: implications for vaccine design and host-pathogen interactions. BMC Microbiol. 10, 173 (2010).
Friedrich, R. et al. Staphylocoagulase is a prototype for the mechanism of cofactor-induced zymogen activation. Nature 425, 535–539 (2003). This paper describes the molecular mechanism for staphylothrombin formation and activity.
Panizzi, P. et al. Fibrinogen substrate recognition by staphylocoagulase (pro)thrombin complexes. J. Biol. Chem. 281, 1179–1187 (2006).
McAdow, M. et al. Preventing Staphylococcus aureus sepsis through the inhibition of its agglutination in blood. PLoS Pathog. 7, e1002307 (2011).
Cheng, A. G. et al. Contribution of coagulases towards Staphylococcus aureus disease and protective immunity. PLoS Pathog. 6, e1001036 (2010). This paper reports that Coa and vWbp are key virulence factors for S. aureus abscess pathogenesis.
McAdow, M. et al. Coagulases as determinants of protective immune responses against Staphylococcus aureus. Infect. Immun. 80, 3389–3398 (2012).
Guggenberger, C., Wolz, C., Morrissey, J. A. & Heesemann, J. Two distinct coagulase-dependent barriers protect Staphylococcus aureus from neutrophils in a three dimensional in vitro infection model. PLoS Pathog. 8, e1002434 (2012).
Ganesh, V. K. et al. A structural model of the Staphylococcus aureus ClfA-fibrinogen interaction opens new avenues for the design of anti-staphylococcal therapeutics. PLoS Pathog. 4, e1000226 (2008).
Thomer, L. et al. N-acetylglucosaminylation of serine-aspartate repeat proteins promotes Staphylococcus aureus blood stream infection. J. Biol. Chem. 289, 3478–3486 (2014).
Foster, T. J., Geoghegan, J. A., Ganesh, V. K. & Höök, M. Adhesion, invasion and evasion: the many functions of the surface proteins of Staphylococcus aureus. Nat. Rev. Microbiol. 12, 49–62 (2014).
Edwards, A. M., Potts, J. R., Josefsson, E. & Massey, R. C. Staphylococcus aureus host cell invasion and virulence in sepsis is facilitated by the multiple repeats within FnBPA. PLoS Pathog. 6, e1000964 (2010).
Heilmann, C. et al. Staphylococcus aureus fibronectin-binding protein (FnBP)-mediated adherence to platelets, and aggregation of platelets induced by FnBPA but not by FnBPB. J. Infect. Dis. 190, 321–329 (2004).
Fitzgerald, J. R. et al. Fibronectin-binding proteins of Staphylococcus aureus mediate activation of human platelets via fibrinogen and fibronectin bridges to integrin GPIIb/IIIa and IgG binding to the FcγRIIa receptor. Mol. Microbiol. 59, 212–230 (2006).
Burke, F. M., McCormack, N., Rindi, S., Speziale, P. & Foster, T. J. Fibronectin-binding protein B variation in Staphylococcus aureus. BMC Microbiol. 10, 160 (2010).
Panther, E. et al. Expression and function of adenosine receptors in human dendritic cells. FASEB J. 15, 1963–1970 (2001).
Csóka, B. et al. A2A adenosine receptors and C/EBPβ are crucially required for IL-10 production by macrophages exposed to Escherichia coli. Blood 110, 2685–2695 (2007).
Thammavongsa, V., Kern, J. W., Missiakas, D. M. & Schneewind, O. Staphylococcus aureus synthesizes adenosine to escape host immune responses. J. Exp. Med. 206, 2417–2427 (2009). The paper describes staphylococcal adenosine synthesis as an immune-evasion mechanism.
Berends, E. T. et al. Nuclease expression by Staphylococcus aureus facilitates escape from neutrophil extracellular traps. J. Innate Immun. 2, 576–586 (2010).
Thammavongsa, V., Missiakas, D. M. & Schneewind, O. Staphylococcus aureus conversion of neutrophil extracellular traps into deoxyadenosine promotes immune cell death. Science 342, 863–866 (2013). This manuscript reports that S. aureus nuclease and AdsA degrade neutrophil NETs to produce deoxyadenosine and kill macrophages.
Forsgren, A. & Nordström, K. Protein A from Staphylococcus aureus: the biological significance of its interaction with IgG. Ann. NY Acad. Sci. 236, 252–266 (1974).
Becker, S., Frankel, M. B., Schneewind, O. & Missiakas, D. M. Release of protein A from the cell wall envelope of Staphylococcus aureus. Proc. Natl Acad. Sci. USA 111, 1574–1579 (2014).
Votintseva, A. A. et al. Prevalence of Staphylococcus aureus protein A (spa) mutants in the community and hospitals in Oxfordshire. BMC Microbiol. 14, 63 (2014).
Shopsin, B. et al. Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J. Clin. Microbiol. 37, 3556–3563 (1999).
Forsgren, A. & Sjöquist, J. Protein A from S. aureus. I. Pseudo-immune reaction with human γ-globulin. J. Immunol. 97, 822–827 (1966).
Sasso, E. H., Silverman, G. J. & Mannik, M. Human IgM molecules that bind staphylococcal protein A contain VHIII H chains. J. Immunol. 142, 2778–2783 (1989).
Forsgren, A. & Quie, P. G. Effects of staphylococcal protein A on heat labile opsonins. J. Immunol. 112, 1177–1180 (1974).
Goodyear, C. S. & Silverman, G. J. Death by a B cell superantigen: in vivo VH-targeted apoptotic supraclonal B cell deletion by a staphylococcal toxin. J. Exp. Med. 197, 1125–1139 (2003).
Falugi, F., Kim, H. K., Missiakas, D. M. & Schneewind, O. The role of protein A in the evasion of host adaptive immune responses by Staphylococcus aureus. mBio 4, e00575–00513 (2013).
Pauli, N. T. et al. Staphylococcus aureus infection induces protein A-mediated immune evasion in humans. J. Exp. Med. 211, 2331–2339 (2014). The paper describes the expansion of V H 3 clonal plasmablast populations in humans with S. aureus infection.
Goodyear, C. S. & Silverman, G. J. Staphylococcal toxin induced preferential and prolonged in vivo deletion of innate-like B lymphocytes. Proc. Natl Acad. Sci. USA 101, 11392–11397 (2004).
Kim, H. K., Cheng, A. G., Kim, H.-Y., Missiakas, D. M. & Schneewind, O. Non-toxigenic protein A vaccine for methicillin-resistant Staphylococcus aureus infections. J. Exp. Med. 207, 1863–1870 (2010). The manuscript describes a vaccine that blocks activity of SpA, a B cell superantigen, to elicit protective immune responses during staphylococcal infection.
Jardetzky, T. S. et al. Three-dimensional structure of a human class II histocompatibility molecule complexed with superantigen. Nature 368, 711–718 (1994).
Fraser, J. D. & Proft, T. The bacterial superantigen and superantigen-like proteins. Immunol. Rev. 225, 226–243 (2008).
Stach, C. S., Herrera, A. & Schlievert, P. M. Staphylococcal superantigens interact with multiple host receptors to cause serious diseases. Immunol. Res. 59, 177–181 (2014).
Choi, Y. et al. Selective expansion of T cells expressing V β 2 in toxic shock syndrome. J. Exp. Med. 172, 981–984 (1990).
Ferry, T. et al. Analysis of superantigenic toxin Vβ T-cell signatures produced during cases of staphylococcal toxic shock syndrome and septic shock. Clin. Microbiol. Infect. 14, 546–554 (2008).
MacDonald, H. R., Lees, R. K., Baschieri, S., Herrmann, T. & Lussow, A. R. Peripheral T-cell reactivity to bacterial superantigens in vivo: the response/anergy paradox. Immunol. Rev. 133, 105–117 (1993).
Ziegler, C. et al. The dynamics of T cells during persistent Staphylococcus aureus infection: from antigen-reactivity to in vivo anergy. EMBO Mol. Med. 3, 652–666 (2011).
Collins, J., Buckling, A. & Massey, R. C. Identification of factors contributing to T-cell toxicity of Staphylococcus aureus clinical isolates. J. Clin. Microbiol. 46, 2112–2114 (2008).
Nakamura, Y. et al. Staphylococcus δ-toxin promotes allergic skin disease by inducing mass cell degranulation. Nature 503, 397–401 (2013). This paper discovers S. aureus δ-toxin as an agent of mast cell degranulation and atopic disease.
Novick, R. P. & Geisinger, E. Quorum sensing in staphylococci. Annu. Rev. Genet. 42, 541–564 (2008).
Rogers, D. E. & Melly, M. A. Speculation on the immunology of staphylococcal infections. Ann. NY Acad. Sci. 128, 274–284 (1965).
Holtfreter, S. et al. Antibody responses in furunculosis patients vaccinated with autologous formalin-killed Staphylococcus aureus. Eur. J. Clin. Microbiol. Infect. Dis. 30, 707–717 (2011).
Weisman, L. E. et al. A randomized study of a monoclonal antibody (pagibaximab) to prevent staphylococcal sepsis. Pediatrics 128, 271–279 (2011).
Fowler, V. G. et al. Effect of an investigational vaccine for preventing Staphylococcus aureus infections after cardiothoracic surgery: a randomized trial. JAMA 309, 1368–1378 (2013).
Shinefield, H. et al. Use of a Staphylococcus aureus conjugate vaccine in patients receiving hemodialysis. N. Engl. J. Med. 346, 491–496 (2002).
Lancefield, R. Current knowledge of type-specific M antigens of group A streptococci. J. Immunol. 89, 307–313 (1962).
Lussow, A. R. & MacDonald, H. R. Differential effects of superantigen-induced “anergy” on priming and effector stages of a T cell-dependent antibody response. Eur. J. Immunol. 24, 445–449 (1994). This report identifies antibodies with broad neutralizing activity for S. aureus β-PFTs.
Rouha, H. et al. Five birds, one stone: neutralization of α-hemolysin and four bi-component leukocidins of Staphylococcus aureus with a single human monoclonal antibody. mAbs 7, 243–254 (2015).
Fitzgerald, J. R. Livestock-associated Staphylococcus aureus: origin, evolution and public health threat. Trends Microbiol. 20, 192–198 (2012).
Lindsay, J. A. Staphylococcus aureus genomics and the impact of horizontal gene transfer. Int. J. Med. Microbiol. 304, 103–109 (2013).
Holt, D. C. et al. A very early-branching Staphylococcus aureus lineage lacking the carotenoid pigment staphyloxanthin. Genome Biol. Evol. 3, 881–895 (2011).
Baker, H. M. et al. Crystal structures of the staphylococcal toxin SSL5 in complex with sialyl Lewis X reveal a conserved binding site that shares common features with viral and bacterial sialic acid binding proteins. J. Mol. Biol. 374, 1298–1308 (2007).
Geisbrecht, B. V., Hamaoka, B. Y., Perman, B., Zemla, A. & Leahy, D. J. The crystal structures of EAP domains from Staphylococcus aureus reveal an unexpected homology to bacterial superantigens. J. Biol. Chem. 280, 17243–17250 (2005).
Lambris, J. D., Ricklin, D. & Geisbrecht, B. V. Complement evasion by human pathogens. Nat. Rev. Microbiol. 6, 132–142 (2008).
Acknowledgements
Work on staphylococcal immune evasion was supported by grants from the US National Institute of Allergy and Infectious Diseases AI038897 (O.S.), AI052747 (O.S.) and AI110937 (D.M.). V.T. acknowledges support from the American Heart Association (PST4590023). We apologize to authors whose work was either not referenced or discussed owing to space constraints.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare a conflict of interest as inventors of patents under licence for commercial development of Staphylococcus aureus vaccines.
Glossary
- Abscesses
-
The pathological product of Staphylococcus aureus infection: the harbouring of a staphylococcal abscess community within a pseudocapsule of fibrin deposits that is surrounded by layers of infiltrating immune cells destroying physiological organ tissue.
- Recurrence
-
The propensity of S. aureus infections to reoccur when surgery and/or antibiotic therapy are initially effective.
- Superantigen
-
Molecules that crosslink B cell receptors (that is, IgM) or T cell receptors and major histocompatibility complexes to trigger lymphocyte proliferation, thereby diverting adaptive immune responses.
- Anaphylatoxin
-
Protein fragments generated during complement activation of C3a and C5a that trigger immune responses via C3a and C5a receptors on immune cells.
- Enterotoxin
-
A staphylococcal superantigen that crosslinks major histocompatibility complex class II molecules and T cell receptors, thereby triggering T cell proliferation, anergy and cytokine storms.
- Opsonization
-
Deposition of complement components on bacterial surfaces to promote recognition, phagocytosis and killing by host phagocytes.
- Fibrinogen
-
An abundant glycoprotein of vertebrates that, when cleaved by thrombin or staphylothombin, self-assembles into fibrin clots.
- FcαRI
-
The IgA Fc receptor, which regulates mucosal immune responses in humans.
- Fcγ domain
-
The portion of antibodies dedicated to C1q complement and Fc-receptor activation.
- Core genome
-
The portion of the genome shared by all members of a bacterial species.
- Leukocidins
-
Bacterial secreted toxins targeting white blood cells (leukocytes) for destruction.
- Dabigatran
-
A small molecule that directly binds and inhibits thrombin as well as staphylothrombin, the complex formed between coagulase or von Willebrand Factor-binding protein and prothrombin.
- Sortase
-
The bacterial transpeptidase responsible for anchoring surface proteins to the cell wall envelope.
- Fibronectin
-
A high molecular weight glycoprotein of the extracellular matrix of vertebrates that associates with integrins on cell surfaces.
- Factor Xa
-
The activated serine protease that cleaves prothrombin to activate the clotting cascade of vertebrates; also known as thrombokinase.
- Fab domains
-
The portions of antibodies dedicated to antigen binding.
- VH3 clan IgM
-
IgM derived from one of three clans of variable heavy chain (VH) genes, the products of which provide the scaffold for the antigen-binding determinants of antibodies.
- Plasmablasts
-
Immature B cells in the blood that secrete antibodies.
- Autologous vaccine
-
A whole-cell killed S. aureus vaccine administered to an infected individual that was derived from the patients isolate.
Rights and permissions
About this article
Cite this article
Thammavongsa, V., Kim, H., Missiakas, D. et al. Staphylococcal manipulation of host immune responses. Nat Rev Microbiol 13, 529–543 (2015). https://doi.org/10.1038/nrmicro3521
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3521
This article is cited by
-
Staphylococcus aureus host interactions and adaptation
Nature Reviews Microbiology (2023)
-
Harnessing the combined effect of antivirulence agent trans-chalcone with bactericidal curcumin against sortase A enzyme to tackle Gram-positive bacterial infections
Folia Microbiologica (2023)
-
The Therapeutic Potential of 4-Methoxy-1-methyl-2-oxopyridine-3-carbamide (MMOXC) Derived from Ricinine on Macrophage Cell Lines Infected with Methicillin-Resistant Strains of Staphylococcus aureus
Applied Biochemistry and Biotechnology (2023)
-
Draft genome sequence of Staphylococcus aureus sequence type 5 SA01 isolated from bloodstream infection and comparative analysis with reference strains
Functional & Integrative Genomics (2023)
-
Staphylococcus aureus populations from the gut and the blood are not distinguished by virulence traits—a critical role of host barrier integrity
Microbiome (2022)