Abstract
The role of the unfolded protein response (UPR) and endoplasmic reticulum (ER) stress in homeostasis of the immune system is incompletely understood. Here we found that dendritic cells (DCs) constitutively activated the UPR sensor IRE-1α and its target, the transcription factor XBP-1, in the absence of ER stress. Loss of XBP-1 in CD11c+ cells led to defects in phenotype, ER homeostasis and antigen presentation by CD8α+ conventional DCs, yet the closely related CD11b+ DCs were unaffected. Whereas the dysregulated ER in XBP-1-deficient DCs resulted from loss of XBP-1 transcriptional activity, the phenotypic and functional defects resulted from regulated IRE-1α-dependent degradation (RIDD) of mRNAs, including those encoding CD18 integrins and components of the major histocompatibility complex (MHC) class I machinery. Thus, a precisely regulated feedback circuit involving IRE-1α and XBP-1 controls the homeostasis of CD8α+ conventional DCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Hetz, C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat. Rev. Mol. Cell Biol. 13, 89–102 (2012).
Ron, D. & Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 8, 519–529 (2007).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Han, D. et al. IRE1α kinase activation modes control alternate endoribonuclease outputs to determine divergent cell fates. Cell 138, 562–575 (2009).
Lee, A.H., Chu, G.C., Iwakoshi, N.N. & Glimcher, L.H. XBP-1 is required for biogenesis of cellular secretory machinery of exocrine glands. EMBO J. 24, 4368–4380 (2005).
Shaffer, A.L. et al. XBP1, downstream of Blimp-1, expands the secretory apparatus and other organelles, and increases protein synthesis in plasma cell differentiation. Immunity 21, 81–93 (2004).
Iwakoshi, N.N., Pypaert, M. & Glimcher, L.H. The transcription factor XBP-1 is essential for the development and survival of dendritic cells. J. Exp. Med. 204, 2267–2275 (2007).
Hollien, J. & Weissman, J.S. Decay of endoplasmic reticulum-localized mRNAs during the unfolded protein response. Science 313, 104–107 (2006).
Hollien, J. et al. Regulated Ire1-dependent decay of messenger RNAs in mammalian cells. J. Cell Biol. 186, 323–331 (2009).
Oikawa, D., Tokuda, M., Hosoda, A. & Iwawaki, T. Identification of a consensus element recognized and cleaved by IRE1α. Nucleic Acids Res. 38, 6265–6273 (2010).
Hur, K.Y. et al. IRE1α activation protects mice against acetaminophen-induced hepatotoxicity. J. Exp. Med. 209, 307–318 (2012).
Upton, J.P. et al. IRE1α cleaves select microRNAs during ER stress to derepress translation of proapoptotic Caspase-2. Science 338, 818–822 (2012).
Kamimura, D. & Bevan, M.J. Endoplasmic reticulum stress regulator XBP-1 contributes to effector CD8+ T cell differentiation during acute infection. J. Immunol. 181, 5433–5441 (2008).
Reimold, A.M. et al. Plasma cell differentiation requires the transcription factor XBP-1. Nature 412, 300–307 (2001).
Iwawaki, T., Akai, R., Kohno, K. & Miura, M. A transgenic mouse model for monitoring endoplasmic reticulum stress. Nat. Med. 10, 98–102 (2003).
Caton, M.L., Smith-Raska, M.R. & Reizis, B. Notch-RBP-J signaling controls the homeostasis of CD8− dendritic cells in the spleen. J. Exp. Med. 204, 1653–1664 (2007).
Lee, A.-H., Scapa, E.F., Cohen, D.E. & Glimcher, L.H. Regulation of hepatic lipogenesis by the transcription factor XBP1. Science 320, 1492–1496 (2008).
Bar-On, L. et al. CX3CR1+ CD8α+ dendritic cells are a steady-state population related to plasmacytoid dendritic cells. Proc. Natl. Acad. Sci. USA 107, 14745–14750 (2010).
Schulz, O. et al. CD40 triggering of heterodimeric IL-12 p70 production by dendritic cells in vivo requires a microbial priming signal. Immunity 13, 453–462 (2000).
Segura, E. & Villadangos, J.A. A modular and combinatorial view of the antigen cross-presentation pathway in dendritic cells. Traffic 12, 1677–1685 (2011).
Sancho, D. et al. Identification of a dendritic cell receptor that couples sensing of necrosis to immunity. Nature 458, 899–903 (2009).
Cebrian, I. et al. Sec22b regulates phagosomal maturation and antigen crosspresentation by dendritic cells. Cell 147, 1355–1368 (2011).
Shoulders, M.D. et al. Stress-independent activation of XBP1s and/or ATF6 reveals three functionally diverse ER proteostasis environments. Cell Rep 3, 1279–1292 (2013).
Acosta-Alvear, D. et al. XBP1 controls diverse cell type- and condition-specific transcriptional regulatory networks. Mol. Cell 27, 53–66 (2007).
Varadarajan, S. et al. Endoplasmic reticulum membrane reorganization is regulated by ionic homeostasis. PLoS ONE 8, e56603 (2013).
Iwawaki, T., Akai, R. & Kohno, K. IRE1α disruption causes histological abnormality of exocrine tissues, increase of blood glucose level, and decrease of serum immunoglobulin level. PLoS ONE 5, e13052 (2010).
Todd, D.J., Lee, A.-H. & Glimcher, L.H. The endoplasmic reticulum stress response in immunity and autoimmunity. Nat. Rev. Immunol. 8, 663–674 (2008).
Satpathy, A.T., Wu, X., Albring, J.C. & Murphy, K.M. Re(de)fining the dendritic cell lineage. Nat. Immunol. 13, 1145–1154 (2012).
Hetz, C. & Glimcher, L.H. Fine-tuning of the unfolded protein response: Assembling the IRE1α interactome. Mol. Cell 35, 551–561 (2009).
McGehee, A.M. et al. XBP-1-deficient plasmablasts show normal protein folding but altered glycosylation and lipid synthesis. J. Immunol. 183, 3690–3699 (2009).
Hu, C.C., Dougan, S.K., McGehee, A.M., Love, J.C. & Ploegh, H.L. XBP-1 regulates signal transduction, transcription factors and bone marrow colonization in B cells. EMBO J. 28, 1624–1636 (2009).
Helft, J. et al. Cross-presenting CD103+ dendritic cells are protected from influenza virus infection. J. Clin. Invest. 122, 4037–4047 (2012).
Luber, C.A. et al. Quantitative proteomics reveals subset-specific viral recognition in dendritic cells. Immunity 32, 279–289 (2010).
den Haan, J.M. & Bevan, M.J. Constitutive versus activation-dependent cross-presentation of immune complexes by CD8+ and CD8− dendritic cells in vivo. J. Exp. Med. 196, 817–827 (2002).
Ahrens, S. et al. F-actin is an evolutionarily conserved damage-associated molecular pattern recognized by DNGR-1, a receptor for dead cells. Immunity 36, 635–645 (2012).
Segura, E., Albiston, A.L., Wicks, I.P., Chai, S.Y. & Villadangos, J.A. Different cross-presentation pathways in steady-state and inflammatory dendritic cells. Proc. Natl. Acad. Sci. USA 106, 20377–20381 (2009).
Zelenay, S. et al. The dendritic cell receptor DNGR-1 controls endocytic handling of necrotic cell antigens to favor cross-priming of CTLs in virus-infected mice. J. Clin. Invest. 122, 1615–1627 (2012).
Guermonprez, P. et al. ER-phagosome fusion defines an MHC class I cross-presentation compartment in dendritic cells. Nature 425, 397–402 (2003).
Nakaya, H.I. et al. Systems biology of vaccination for seasonal influenza in humans. Nat. Immunol. 12, 786–795 (2011).
Sha, H. et al. The IRE1alpha-XBP1 pathway of the unfolded protein response is required for adipogenesis. Cell Metab. 9, 556–564 (2009).
Martinon, F., Chen, X., Lee, A.-H. & Glimcher, L.H. TLR activation of the transcription factor XBP1 regulates innate immune responses in macrophages. Nat. Immunol. 11, 411–418 (2010).
Lee, A.-H., Scapa, E.F., Cohen, D.E. & Glimcher, L.H. Regulation of hepatic lipogenesis by the transcription factor XBP1. Science 320, 1492–1496 (2008).
Acknowledgements
We thank L. Glimcher (Cornell University) for Xbp1fl/fl mice; B. Reizis (Columbia University) for Itgax-Cre mice; B. Malissen (Aix Marseille University) for H-Y mice; J. Magalhaes (Institut Curie) for advice and protocols; D. Jankovic and A. Sher (National Institute of Allergy and Infectious Diseases) for T. gondii extracts; C. Reis e Sousa (Cancer Research UK) for bm1 T OVA cells; the VIB Microarray Facility for doing the microarray experiments; the IRC Microscopy Core Facility and, more specifically, A. Kremers, S. Lippens and C. Guerin for help with the imaging of XBP-1-deficient CD8α+ cDCs by focused ion beam–scanning electron microscopy. Supported by the European Research Council (B.N.L.), the European Union Seventh Framework Programme (B.N.L.), the Fonds Wetenschappelijk Onderzoek Vlaanderen program (B.N.L. and S.J.), Ghent University (B.N.L. and S.J.), Marie Curie Actions (F.O. and E.H.), The Federation of European Biochemical Societies (F.O.) and the Agentschap voor Innovatie door Wetenschap en Techniek (S.J.T.).
Author information
Authors and Affiliations
Contributions
F.O., S.J. and B.N.L. designed the research; F.O., S.J.T., E.H. and P.P. did the experiments; F.O. analyzed the results; Y.S. and L.M. helped with microarray analysis; R.D.R. did transmission electron microscopy, E.P. helped with microscopy analysis; J.V. and I.D. provided technical assistance; T.I. provided reagents; and F.O., S.J. and B.N.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Activation of the IRE-1α pathway in peripheral DCs.
Expression of VenusFP in (a) lung tissue or (b) lung draining lymph node (Lung dr. LN) from WT and ERAI reporter mice. (c and d) Histograms depict the VenusFP expression in B cells (CD19+, MHCII+) T cells (CD3e+) and DC (Lin-, MHCII+, CD11c+) in control animals (grey) and ERAI mice (blue) in lung (n=3) or lung dr. LN (n=2) and is representative of three independent experiments. (e) Schematic representation of the generation of the mixed BM chimeras.
Supplementary Figure 2 Detail of aberrant ER morphology in XBP-1-deficient CD8α+ cDCs.
Electron micrographs of FACS-sorted CD8α+ cDCs derived from DC-XBP-1Δ or control mice. Scale bars, 250nm. Data is representative of one experiment.
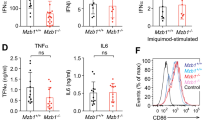
Supplementary Figure 3 CD8α+ cDCs from DC–XBP-1Δ mice are as efficient as CD8α+ cDCs from their control littermates in secreting IL-12.
IL-12-p70 concentration in serum (left panel) or splenic cell supernatant (right panel) from DC-XBP-1Δ mice or control littermates after injection with STAg. IL-12 concentration was quantified by ELISA. Bars represent mean ± SEM of two independent experiments. p=0,095 (Mann Whitney U test, two-sided).
Supplementary Figure 4 DCs from DC–XBP-1Δ mice sustain CD4+ T cell proliferation, and early modules of cross-presentation are normal in XBP-1-deficient CD8α+ cDCs.
FACS-sorted CD11b+ cDCs (a) or purified CD8α+ cDCs (b) from DC-XBP-1Δ mice or control littermates were cultured with CFSE-labeled OVA specific TCR Tg CD4+ OT-II cells in presence of OVA peptide and soluble OVA. Cell counts were measured on day three. Plots depict mean ± SEM of two independent experiments. (c) Phagocytosis of 1μm polystyrene microspheres conjugated to OVA after labeling of remaining microspheres with an Alexa 488-conjugated antibody. Right gate in dot plots depicts frequency of cells with surface-bound beads and left gate depicts frequency of cells with internalized particles. Data are representative of 2 independent experiments. (d) Cells were loaded with a cytosolic FRET-sensitive substrate of β-lactamase (βlac) and incubated in absence of presence of βlac. After 3 h, change in fluorescence in gated CD8α+ cDCs was measured as readout of βlac export from the endocytic pathway into the cytosol. Data are representative of 2 independent experiments.
Supplementary Figure 5 Loss of CD18 in XBP-1-deficient CD8α+ cDCs results in decreased expression of CD11c and CD11a.
(a) CD11a (LFA-1) expression in splenic CD11b+ and CD8α+ cDCs in DC-XBP-1Δ mice and control littermates. (b) Table of reported UPR target genes with fold induction (FC) below twofold in XBP-1 deficient versus sufficient CD8α+ cDCs 4,24,25,44.
Supplementary Figure 6 Schematic representation of the IRE-1α cleavage sites with mRNA secondary structure prediction.
Potential IRE-1α splicing motifs were extracted for Itgb2 (CD18), Ergic3 and Tapbp. Secondary structure prediction was performed using three different programs: Mfold (depicted), CentroidFold and RNAFold. Sequence motifs where at least two of the three programs agreed on the predicted secondary structure were retained, and further filtering was performed by retaining those motifs where the cleavage site was situated in the loop structure. Arrows indicate the splicing site.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 4404 kb)
Aberrant ER morphology in XBP-1-deficient CD8α+ cDCs.
3D-imaging of a XBP deficient CD8α+ cDC using FIB-SEM, first showing the raw data and aberrant ER, which is partially reconstructed in green to show the increased complexity of the ER in these cells. (AVI 31681 kb)
Rights and permissions
About this article
Cite this article
Osorio, F., Tavernier, S., Hoffmann, E. et al. The unfolded-protein-response sensor IRE-1α regulates the function of CD8α+ dendritic cells. Nat Immunol 15, 248–257 (2014). https://doi.org/10.1038/ni.2808
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2808
This article is cited by
-
Immunometabolism: a new dimension in immunotherapy resistance
Frontiers of Medicine (2023)
-
Adipocyte IRE1α promotes PGC1α mRNA decay and restrains adaptive thermogenesis
Nature Metabolism (2022)
-
PERK is a critical metabolic hub for immunosuppressive function in macrophages
Nature Immunology (2022)
-
Saponin-based adjuvant-induced dendritic cell cross-presentation is dependent on PERK activation
Cellular and Molecular Life Sciences (2022)
-
Evolution and function of the epithelial cell-specific ER stress sensor IRE1β
Mucosal Immunology (2021)