Abstract
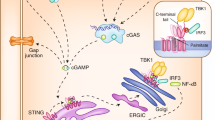
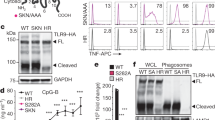
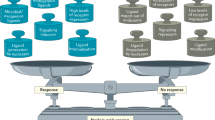
The activation of innate immune responses by nucleic acids is crucial to protective and pathological immunities and is mediated by the transmembrane Toll-like receptors (TLRs) and cytosolic receptors1,2. However, it remains unknown whether a mechanism exists that integrates these nucleic-acid-sensing systems. Here we show that high-mobility group box (HMGB) proteins 1, 2 and 3 function as universal sentinels for nucleic acids. HMGBs bind to all immunogenic nucleic acids examined with a correlation between affinity and immunogenic potential. Hmgb1-/- and Hmgb2-/- mouse cells are defective in type-I interferon and inflammatory cytokine induction by DNA or RNA targeted to activate the cytosolic nucleic-acid-sensing receptors; cells in which the expression of all three HMGBs is suppressed show a more profound defect, accompanied by impaired activation of the transcription factors interferon regulatory factor 3 (IRF3) and nuclear factor (NF)-κB. The absence of HMGBs also severely impairs the activation of TLR3, TLR7 and TLR9 by their cognate nucleic acids. Our results therefore indicate a hierarchy in the nucleic-acid-mediated activation of immune responses, wherein the selective activation of nucleic-acid-sensing receptors is contingent on the more promiscuous sensing of nucleic acids by HMGBs. These findings may have implications for understanding the evolution of the innate immune system and for the treatment of immunological disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Akira, S., Uematsu, S. & Takeuchi, O. Pathogen recognition and innate immunity. Cell 124, 783–801 (2006)
Medzhitov, R. Recognition of microorganisms and activation of the immune response. Nature 449, 819–826 (2007)
Hornung, V. et al. Sequence-specific potent induction of IFN-α by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nature Med. 11, 263–270 (2005)
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006)
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nature Immunol. 5, 730–737 (2004)
Ishii, K. J. et al. TANK-binding kinase-1 delineates innate and adaptive immune responses to DNA vaccines. Nature 451, 725–729 (2008)
Schroder, K., Muruve, D. A. & Tschopp, J. Innate immunity: cytoplasmic DNA sensing by the AIM2 inflammasome. Curr. Biol. 19, R262–R265 (2009)
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–505 (2007)
Choi, M. K. et al. A selective contribution of the RIG-I-like receptor pathway to type I interferon responses activated by cytosolic DNA. Proc. Natl Acad. Sci. USA advance online publication 10.1073/pnas.0910983106 (1 October 2009)
Chiu, Y. H., Macmillan, J. B. & Chen, Z. J. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009)
Ablasser, A. et al. RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III–transcribed RNA intermediate. Nature Immunol. 10, 1065–1072 (2009)
Ishii, K. J. et al. A Toll-like receptor-independent antiviral response induced by double-stranded B-form DNA. Nature Immunol. 7, 40–48 (2006)
Bianchi, M. E. & Manfredi, A. A. High-mobility group box 1 (HMGB1) protein at the crossroads between innate and adaptive immunity. Immunol. Rev. 220, 35–46 (2007)
Scaffidi, P., Misteli, T. & Bianchi, M. E. Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature 418, 191–195 (2002)
Apetoh, L. et al. The interaction between HMGB1 and TLR4 dictates the outcome of anticancer chemotherapy and radiotherapy. Immunol. Rev. 220, 47–59 (2007)
Ivanov, S. et al. A novel role for HMGB1 in TLR9-mediated inflammatory responses to CpG-DNA. Blood 110, 1970–1981 (2007)
Kazama, H. et al. Induction of immunological tolerance by apoptotic cells requires caspase-dependent oxidation of high-mobility group box-1 protein. Immunity 29, 21–32 (2008)
Tian, J. et al. Toll-like receptor 9-dependent activation by DNA-containing immune complexes is mediated by HMGB1 and RAGE. Nature Immunol. 8, 487–496 (2007)
Krieg, A. M. et al. CpG motifs in bacterial DNA trigger direct B-cell activation. Nature 374, 546–549 (1995)
Klinman, D. M. Immunotherapeutic uses of CpG oligodeoxynucleotides. Nature Rev. Immunol. 4, 249–258 (2004)
Haas, T. et al. The DNA sugar backbone 2′ deoxyribose determines Toll-like receptor 9 activation. Immunity 28, 315–323 (2008)
Nemeth, M. J. et al. Hmgb3: an HMG-box family member expressed in primitive hematopoietic cells that inhibits myeloid and B-cell differentiation. Blood 102, 1298–1306 (2003)
Vaccari, T., Beltrame, M., Ferrari, S. & Bianchi, M. E. Hmg4, a new member of the Hmg1/2 gene family. Genomics 49, 247–252 (1998)
Stetson, D. B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006)
Pichlmair, A. & Reis e Sousa, C. Innate recognition of viruses. Immunity 27, 370–383 (2007)
Hornung, V. et al. 5′-Triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006)
Paladino, P., Cummings, D. T., Noyce, R. S. & Mossman, K. L. The IFN-independent response to virus particle entry provides a first line of antiviral defense that is independent of TLRs and retinoic acid-inducible gene I. J. Immunol. 177, 8008–8016 (2006)
Grasser, K. D., Launholt, D. & Grasser, M. High mobility group proteins of the plant HMGB family: dynamic chromatin modulators. Biochim. Biophys. Acta 1769, 346–357 (2007)
Kampmann, T., Mueller, D. S., Mark, A. E., Young, P. R. & Kobe, B. The role of histidine residues in low-pH-mediated viral membrane fusion. Structure 14, 1481–1487 (2006)
Weissenhorn, W., Hinz, A. & Gaudin, Y. Virus membrane fusion. FEBS Lett. 581, 2150–2155 (2007)
Calogero, S. et al. The lack of chromosomal protein Hmg1 does not disrupt cell growth but causes lethal hypoglycaemia in newborn mice. Nature Genet. 22, 276–280 (1999)
Ronfani, L. et al. Reduced fertility and spermatogenesis defects in mice lacking chromosomal protein Hmgb2. Development 128, 1265–1273 (2001)
Honda, K. et al. Spatiotemporal regulation of MyD88–IRF-7 signalling for robust type-I interferon induction. Nature 434, 1035–1040 (2005)
Acknowledgements
We thank C. Reis e Sousa and J. Rehwinkel for 5′-triphosphate RNA;, Y. Kawaguchi for HSV-1 DNA; A. Katoh and M. Kidokoro for the vaccinia virus (MO) genome for ligand stimulation; J. Vilcek, R. Medzhitov, R. Schekman, J. V. Ravetch and H. Ohno for invaluable advice; and R. Takeda, M. Shishido, Y. Nakaima and T. Oh for technical assistance. This work was supported in part by a Grant-in-Aid for Scientific Research on Priority Areas ‘Integrative Research Toward the Conquest of Cancer,’ Grant-in-Aid for Scientific Research (A), and Global Center of Excellence Program ‘Integrative Life Science Based on the Study of Biosignaling Mechanisms’ from the Ministry of Education, Culture, Sports, Science, and Technology of Japan, and by the Korea Science and Engineering Foundation Grant. T.B. and Z.W. are research fellows of the Japan Society for the Promotion of Science.
Author Contributions H.Y., T.B., Z.W. and M.C. designed, performed and analysed most of the experiments in this study. T. Kawamura and T. Kodama collected data and performed the mass spectrometry analysis. H.N. performed experiments on RAW264.7 cells. M.N. performed the fluorescence microscopy analysis. Y.L. and R.K. cloned the constructs. S.H. assisted with the virus infection experiments. A.S., L.R. and M.B. provided precious materials and invaluable advice. K.H., T. Tamura and T. Taniguchi provided overall coordination with respect to conception, design and supervision of the study. H.Y., T.B., D.S., T. Tamura and T. Taniguchi wrote the manuscript with comments from the co-authors.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References and Supplementary Figures 1-7 with Legends. (PDF 1411 kb)
Rights and permissions
About this article
Cite this article
Yanai, H., Ban, T., Wang, Z. et al. HMGB proteins function as universal sentinels for nucleic-acid-mediated innate immune responses . Nature 462, 99–103 (2009). https://doi.org/10.1038/nature08512
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08512
This article is cited by
-
Macrophage-derived exosomal HMGB3 regulates silica-induced pulmonary inflammation by promoting M1 macrophage polarization and recruitment
Particle and Fibre Toxicology (2024)
-
Proteomic study of left ventricle and cortex in rats after myocardial infarction
Scientific Reports (2024)
-
The multifunctional protein HMGB1: 50 years of discovery
Nature Reviews Immunology (2023)
-
Role of alarmins in poststroke inflammation and neuronal repair
Seminars in Immunopathology (2023)
-
CXCL12/CXCR4 Axis is Involved in the Recruitment of NK Cells by HMGB1 Contributing to Persistent Airway Inflammation and AHR During the Late Stage of RSV Infection
Journal of Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.